More Comprehensive and Updated Coverage
At the design level, QuarStar Human All Exon Probes 4.0 (Heredity Version) selects core coding exonic regions from the latest versions of mainstream databases, achieving nearly 99.5% target design coverage.
Excluding the CNV backbone, the total target region is 41.05 Mb, with a probe-covered region of 45.58 Mb, resulting an optimized bait-to-target ratio of 1.11, ensuring comprehensive coverage while minimizing unnecessary probe redundancy and sequencing cost.
Table 1. QuarStar Human All Exon Probes 4.0 (Heredity Version) Database and Coverage
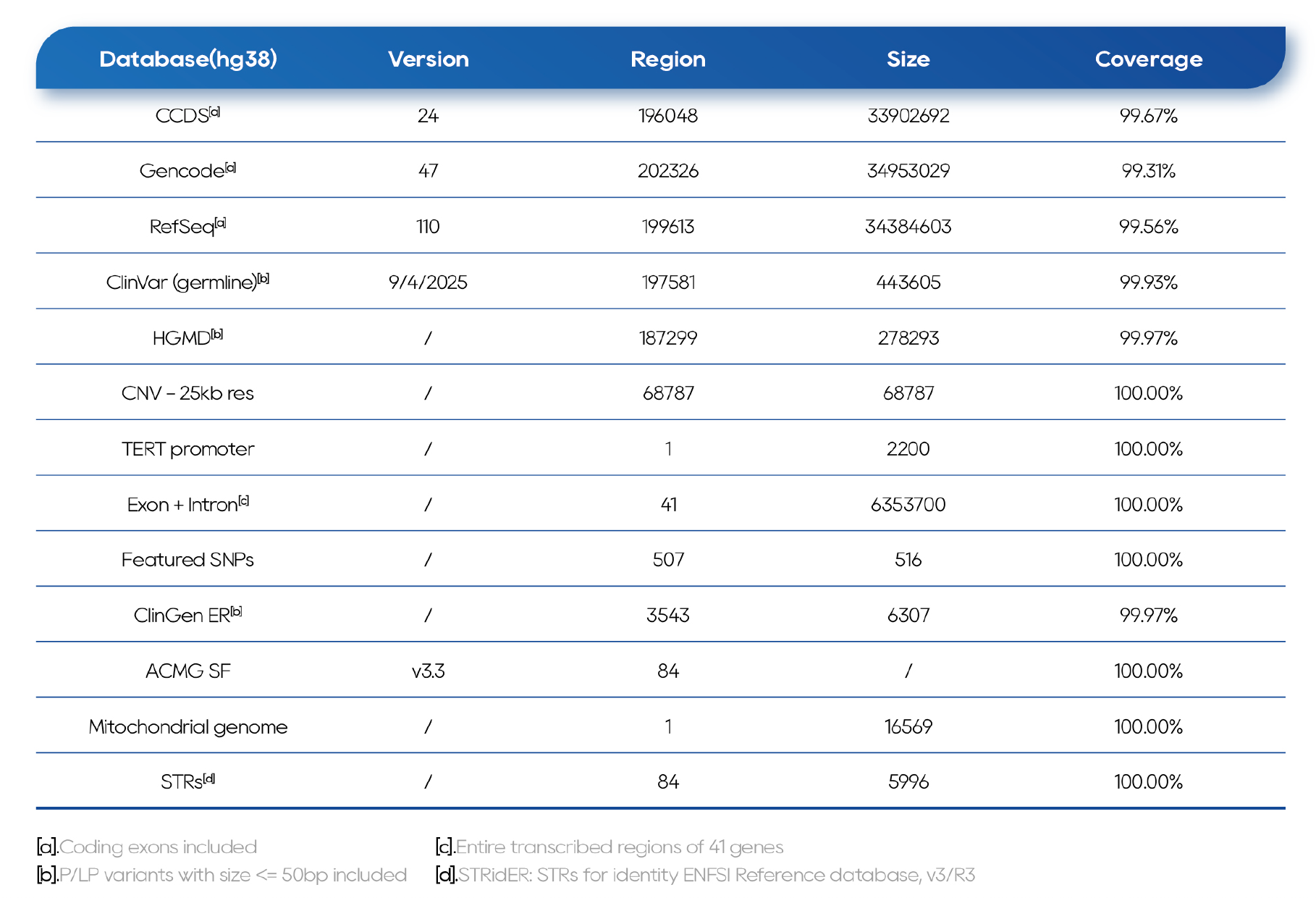
Outstanding Performance
Leveraging its proprietary high-throughput synthesis platform, Dynegene enables large-scale probe production together with iterative optimization of multiple probe design versions. AI-driven probe design further improves on-target rate and target coverage. In addition, a non-polarization boosting strategy combined with an unbiased probe amplification process ensures exceptionally high coverage uniformity. Furthermore, a unique double-stranded multi-level probe biotin labeling technology provides significantly higher labeling efficiency than conventional column-synthesized probe end-labeling methods, ensuring sufficient sequencing depth even for low-input samples and challenging variant regions.
/ Industry-Leading GC Bias Control
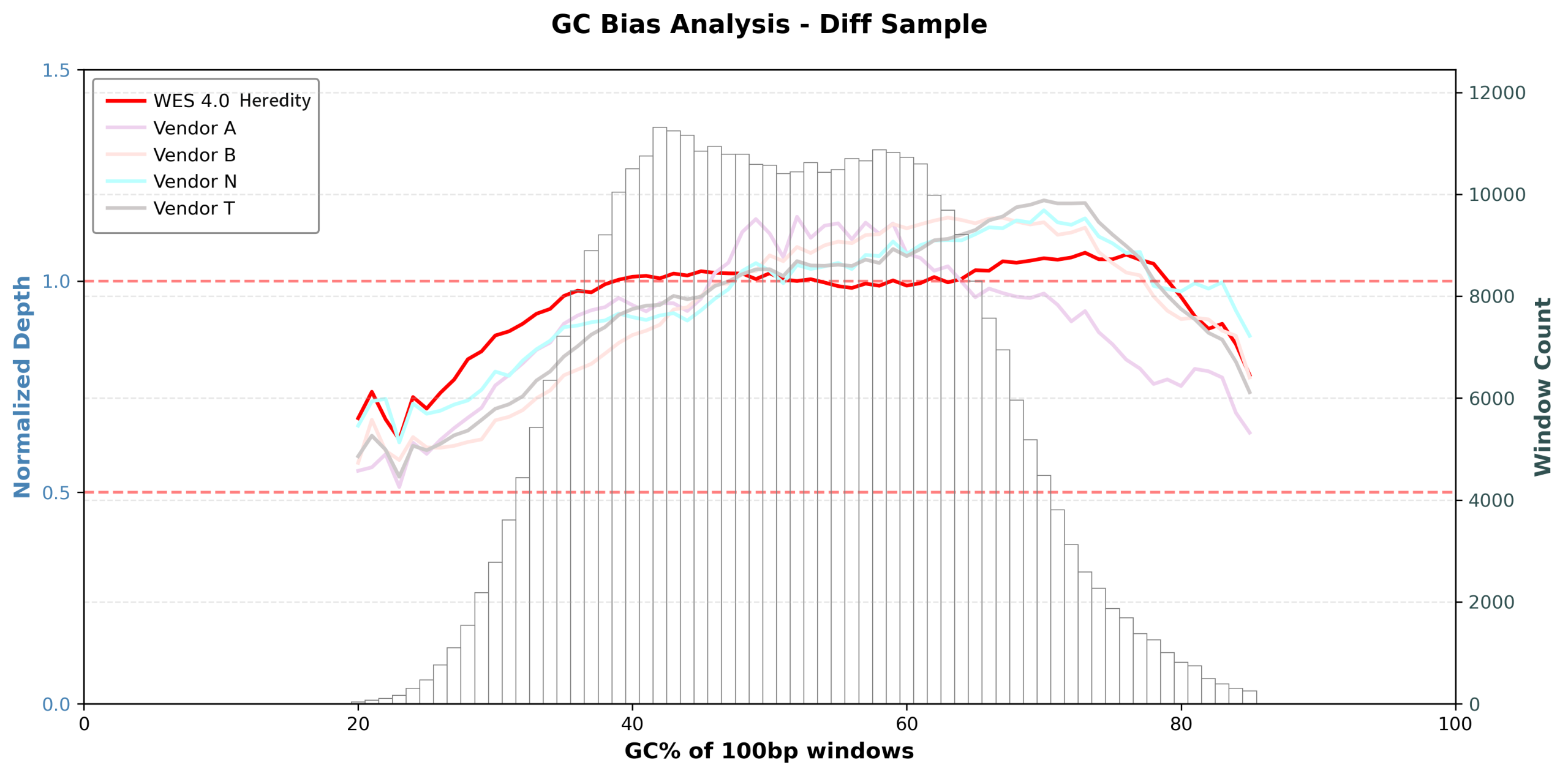
Figure 1: GC Bias Analysis
Regions with GC content from 18% to 86% maintain sequencing depth above 0.5× of the mean depth, while regions between 28% and 82% GC remain within 0.9–1.1× of the mean depth. Within the typical genomic GC range (35%–65%), sequencing depth is approximately equal to the mean depth.
As shown in Figure 1, sequencing depth remains stable across a wide GC content range, which is significantly superior to mainstream domestic and international competitors.
/ Higher On-Target Rate at High Coverage

Figure 2: Core Performance Metrics
· Coverage > 99.5% · On-target rate ≈ 90%
· 0.2× uniformity > 99% · 0.5× uniformity ≈ 93%
· Consistent performance under single-plex and multiplex (12-plex) hybridization conditions
/ Precise Differential Depth Design

Figure 3: Differential Depth Statistics
Based on regional characteristics and detection complexity, differentiated sequencing depth is implemented to maximize data utilization.
· Single-plex: 8 Gb/sample
· 12-plex: 4 Gb/sample
· Effective depth of core exonic regions reaches 11.5× per Gb
(Data includes optional CNV and mitochondrial targets.)
/ Exceptional Uniformity

Figure 4: Fold-80 Statistics
· Core exon uniformity (0.5×) reaches 95.49%
· Fold-80 as low as 1.21
· Overall Fold-80 ranges between 1.3–1.4 under both single-plex and multiplex conditions
/ Superior Variant Detection Capability

Figure 5: F1 Score Statistics
Using the NA12878 reference standard library for capture sequencing:
· SNP F1 score > 99%
· Indel F1 score > 93.5%
Comprehensive Target Content
a)More Comprehensive Target Content
Based on the latest expert consensus, guideline documents, and published literature, QuarStar Human All Exon Probes 4.0 (Heredity Version) covers more than 40 high-frequency complex variant genes (including CNVs and large insertions/deletions) for full transcript coverage, aiming to improve detection rates of complex variants and rare diseases.
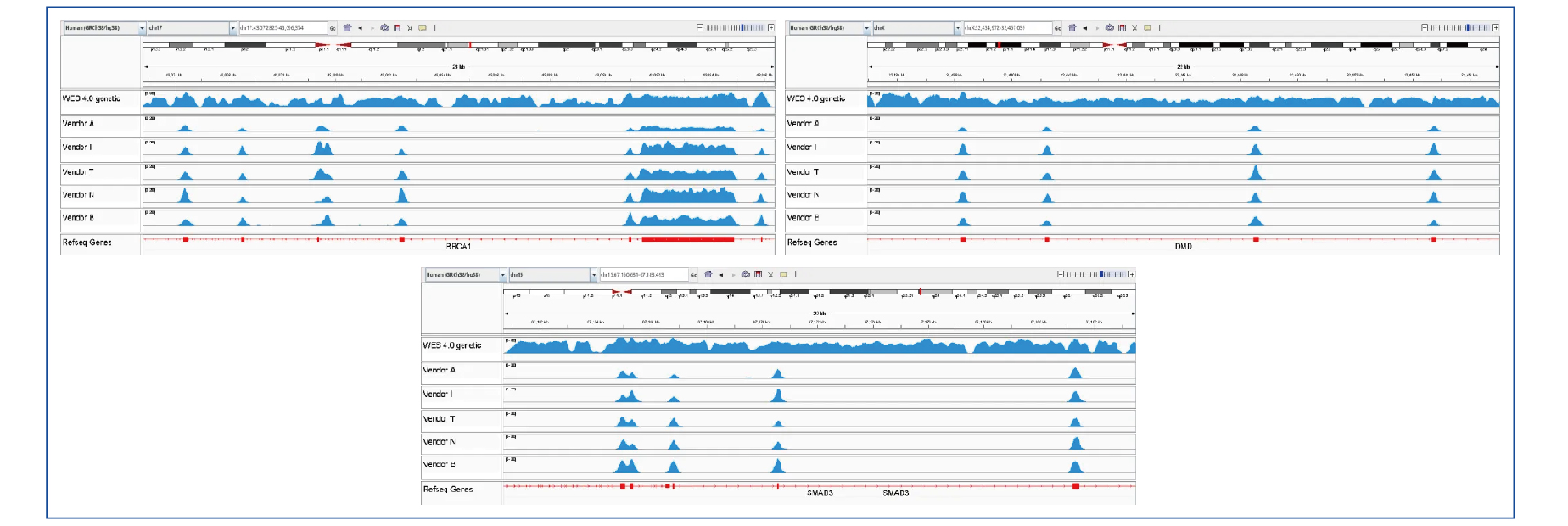
Figure 6: BRCA, DMD, SMAD3 Gene Coverage Comparison
b)Full-length mitochondrial genome coverage reaches 100%, with 0.5× uniformity also at 100% after deduplication.
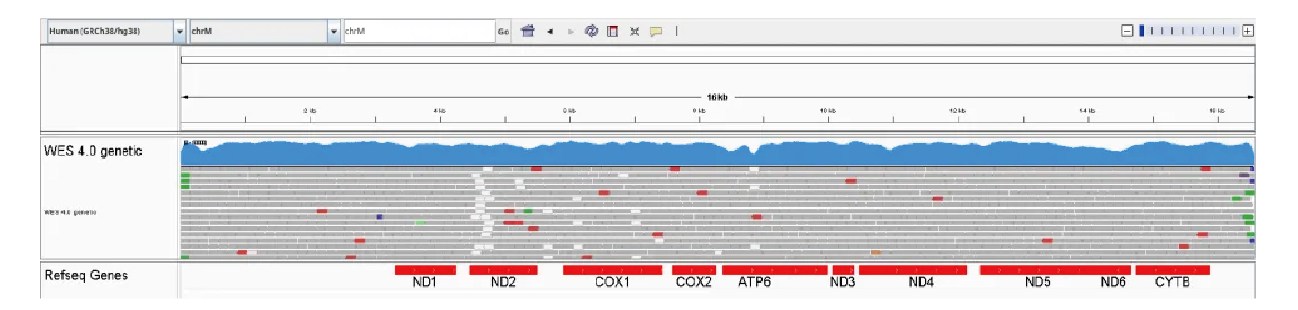
Figure 7: Full Mitochondrial Genome Coverage
The panel also includes ancestry markers, pharmacogenomic loci (e.g., CYP2C19, CYP2D6, TPMT), and sample tracking SNPs to support diverse analyses.
Enhanced Detection of Challenging Regions
a)TERT Promoter Coverage
Sequencing depth of the TERT promoter exceeds 250×, surpassing panel average, with highly uniform coverage.
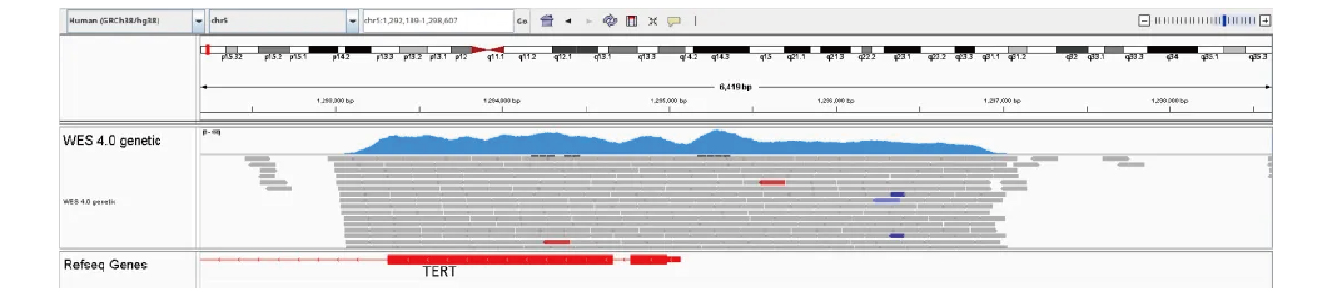
Figure 8: TERT Promoter Hotspot Coverage
b)Improved STR Region Capture
Highly repetitive STR regions are captured with superior performance. For example, MONO-27 and NR-21 repeats (27 T and 21 A repeats) show significantly better coverage than competitor products.
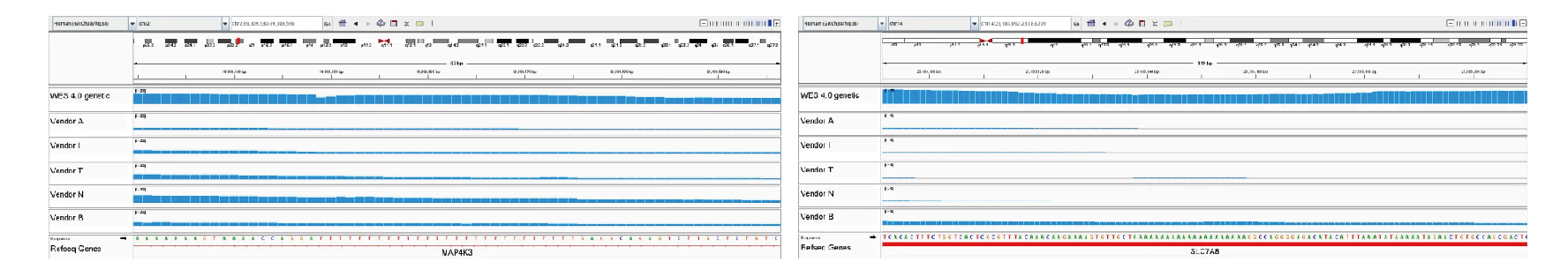
Figure 9: STR Loci Coverage Comparison
c)Improved Capture of High GC Regions
In extremely high-GC regions, such as the GCGR gene and MMP17 exon 1, Dynegene demonstrates significantly improved coverage over competitors.
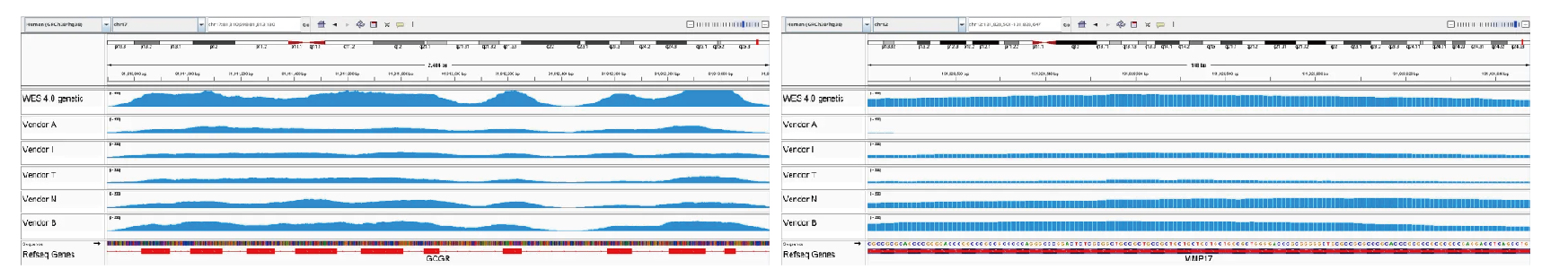
Figure 10: GC-Rich Region Coverage Comparison
One-Stop CNV Detection
/ Ultra-High Uniformity Enables Reliable CNV Detection

Figure 11: Core Exon Uniformity and Fold-80 Parameters
The extremely high coverage uniformity and low GC bias of QuarStar Human All Exon Probes 4.0 (Heredity Version) provide stable read-depth signals across target regions, enabling reliable CNV detection from WES data. This performance may reduce the need for additional baseline reference construction, simplifying the CNV analysis workflow and improving analysis efficiency and stability.
/ High-Density CNV Backbone Probes for Robust Detection
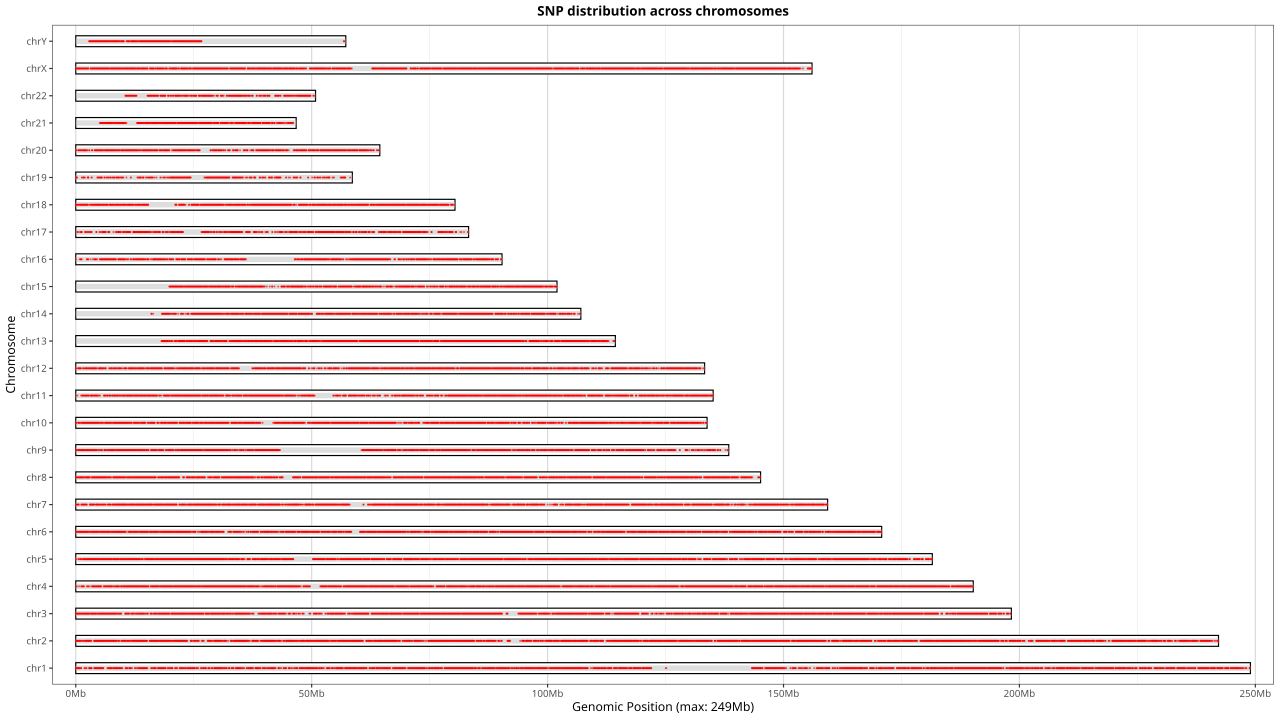
Figure 12: Genome-Wide Coverage of Backbone Probes
Heterozygous SNP loci with stable copy numbers and high MAF were carefully selected across the genome. Loci deviating from reference populations were excluded using Hardy–Weinberg equilibrium testing, and regions in centromeres, telomeres, highly homologous sequences (e.g., pseudogenes), and highly polymorphic regions (e.g., HLA) were removed.
Over 68,000 SNP loci were included, covering ~8.25 Mb of genomic regions, evenly distributed across intergenic and intronic regions to achieve detection resolution up to 25 kb.
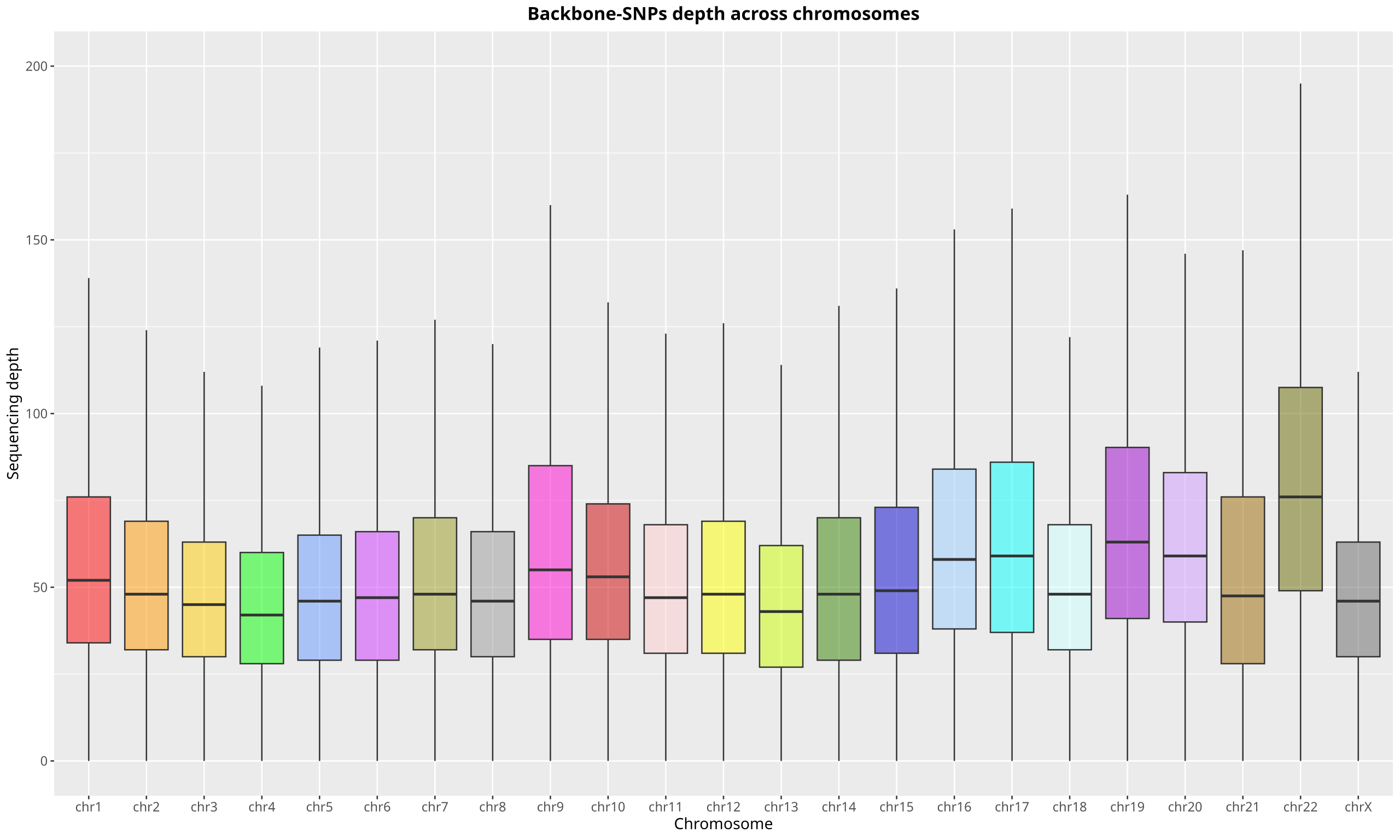
Figure 13: Chromosomal Coverage Depth of SNP Backbone Probes
Using the NA12878 standard sample for hybrid capture sequencing, analysis of SNP site depth across chromosomes (excluding chrY) shows a highly balanced median depth, fully ensuring the backbone regions serve as stable internal references.
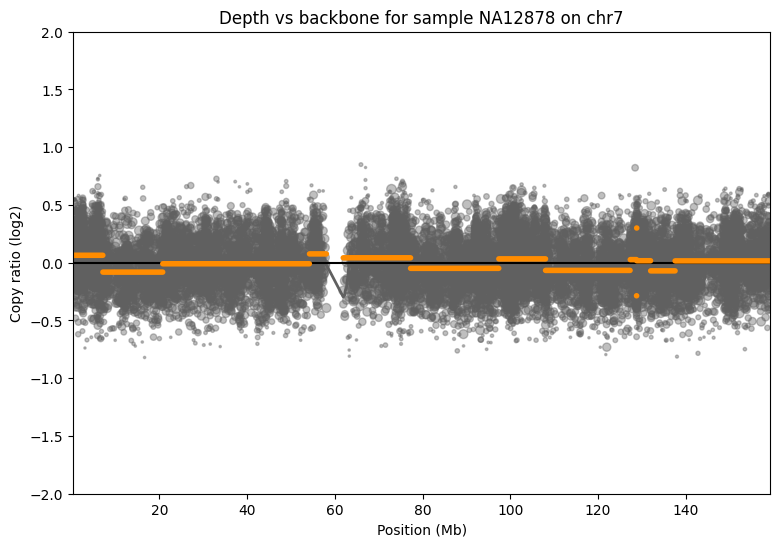
Figure 14: CNV Analysis of Chromosome 7
Analysis using CNVkit (v0.9.12) showed CNV detection based on backbone probes is highly feasible and accurate.
 NGSHybridization Capture Probe NGS custom probes MRD custom probes QuarStar Human MethylCap Panel QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 QuarStar Pan-Cancer Panel 1000-Gene QuarStar HLA Panel QuarXeq HRD panel QuarXeq Mitochondrial Probes Whole Exome Sequencing Probes QuarStar Human All Exon Probes 4.0 (Heredity) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 1.0 (RNA) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Dynegene Adapter Family Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus3 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Blocker Family QuarHyb DNA Reagent Kit Pro(Methyl) Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System
NGSHybridization Capture Probe NGS custom probes MRD custom probes QuarStar Human MethylCap Panel QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 QuarStar Pan-Cancer Panel 1000-Gene QuarStar HLA Panel QuarXeq HRD panel QuarXeq Mitochondrial Probes Whole Exome Sequencing Probes QuarStar Human All Exon Probes 4.0 (Heredity) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 1.0 (RNA) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Dynegene Adapter Family Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus3 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Blocker Family QuarHyb DNA Reagent Kit Pro(Methyl) Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System Primers and Probes
Primers and Probes RNA SynthesissgRNA miRNA siRNA
RNA SynthesissgRNA miRNA siRNA



 Gene
Gene Oligo Pools
Oligo Pools CRISPR sgRNA Library
CRISPR sgRNA Library Antibody Library
Antibody Library Variant Library
Variant Library


















 Tel: 400-017-9077
Tel: 400-017-9077 Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai
Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai Email:
Email: 







