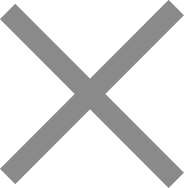
Oligo pools have become a foundational synthesis tool in modern genomics, synthetic biology, CRISPR functional screening, and targeted sequencing. For research teams that need thousands to millions of precisely defined sequences in parallel, oligo pool synthesis offers a scalable and cost-effective path that individual oligo ordering simply cannot match.

At Dynegene, oligo pool synthesis is supported by a next-generation ultra-high-throughput DNA microarray synthesis platform that delivers oligos up to 350 nt in length and up to 4.35 million unique sequences on a single chip — with high accuracy and uniformity as core design requirements, not afterthoughts. These capabilities make oligo pools suitable for a wide spectrum of research and industrial applications, from CRISPR sgRNA library construction and variant library generation to target enrichment probe development, MPRA, FISH, spatial transcriptomics, synthetic biology, and DNA data storage.
What Are Oligo Pools
Oligo pools are highly complex mixtures of individually designed oligonucleotide sequences synthesized simultaneously on a DNA microarray platform. Unlike conventional column-based oligo synthesis — where each sequence occupies a dedicated synthesis column and is delivered as an individual tube — oligo pool synthesis produces thousands to millions of distinct sequences in a single chip run, all delivered together as a pooled mixture.
The key distinction is that each oligo in the pool is sequence-defined by design, not generated randomly. This separates oligo pools from error-prone PCR mutagenesis approaches, which introduce mutations stochastically and without sequence control. Every member of a well-designed oligo pool can be traced back to an explicit design decision, which makes the resulting libraries interpretable in a way that random mutagenesis libraries are not.
This format is especially valuable when experiments depend on a large parallelized design space — CRISPR guide libraries that must cover every exon of a gene family, saturation mutagenesis libraries that test every amino acid at every position, or hybridization probe sets that tile thousands of genomic regions with defined spacing.
Why Oligo Pools Matter for Modern Research
High-throughput biology increasingly depends on scale. A genome-wide CRISPR screen requires guides targeting 18,000 to 20,000 human protein-coding genes, with 3 to 6 guides per gene — that is 60,000 to 120,000 unique sequences for a single experiment. A comprehensive deep mutational scanning study of an antibody CDR region may require 2,000 to 10,000 single-mutant and combinatorial variants. Neither experiment is operationally or economically feasible through one-by-one synthesis.
Oligo pools address this at the synthesis layer:
- Cost efficiency: Cost per designed sequence drops dramatically as library complexity increases, making large-scale library construction accessible to standard research budgets
- Speed: The entire library is synthesized in a single production run rather than assembled from hundreds of individual orders with varying lead times
- Design precision: Every sequence is synthesized to specification, enabling libraries where representation and composition can be predicted and verified
- Downstream compatibility: Pooled oligos are directly compatible with PCR amplification, restriction enzyme cloning, Golden Gate Assembly, Gibson Assembly, lentiviral packaging, and hybridization capture workflows
For teams working across functional genomics, NGS panel development, and synthetic biology, oligo pools are often the rate-limiting step that, when optimized, accelerates every downstream experiment.
Core Applications of Oligo Pools
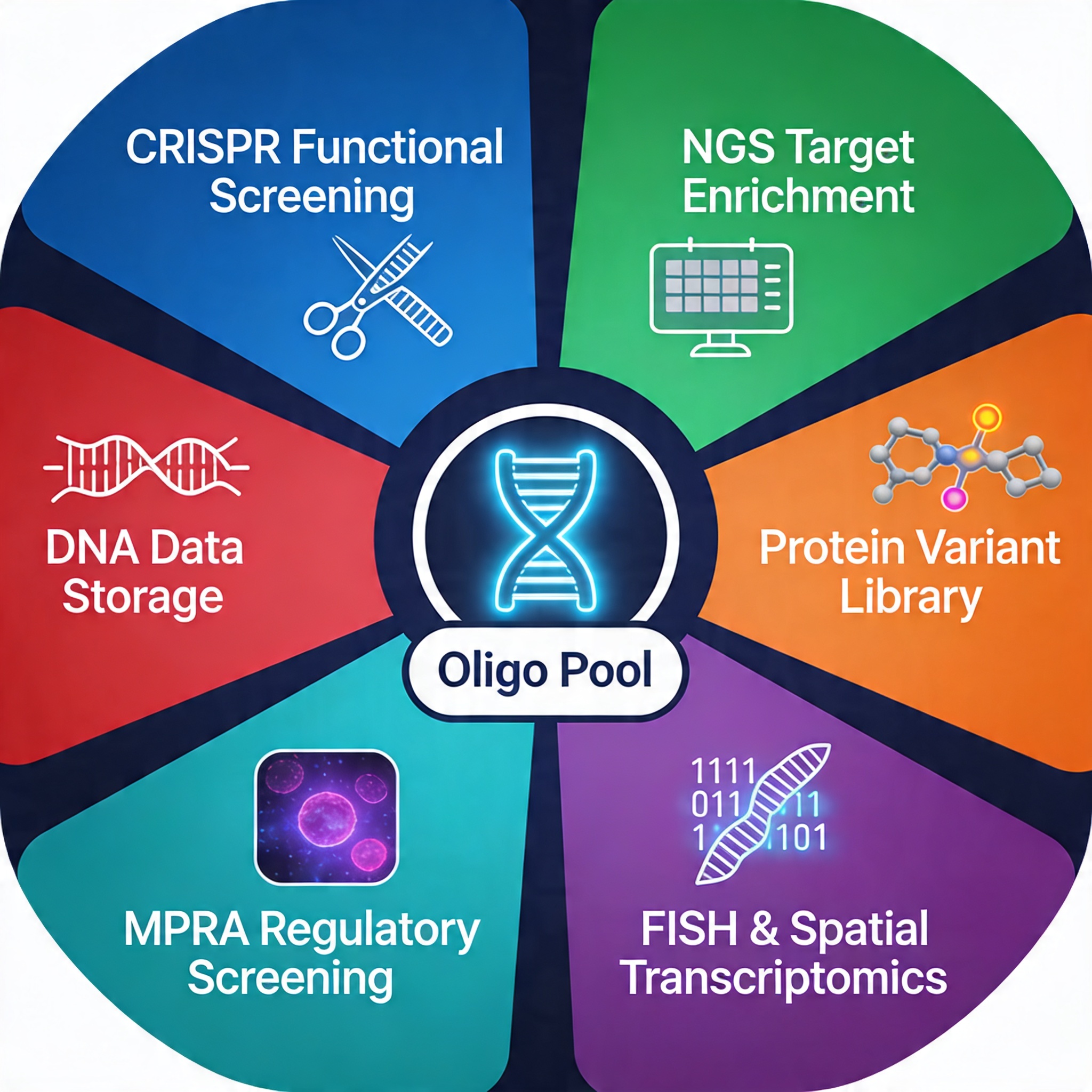
CRISPR sgRNA Library Construction
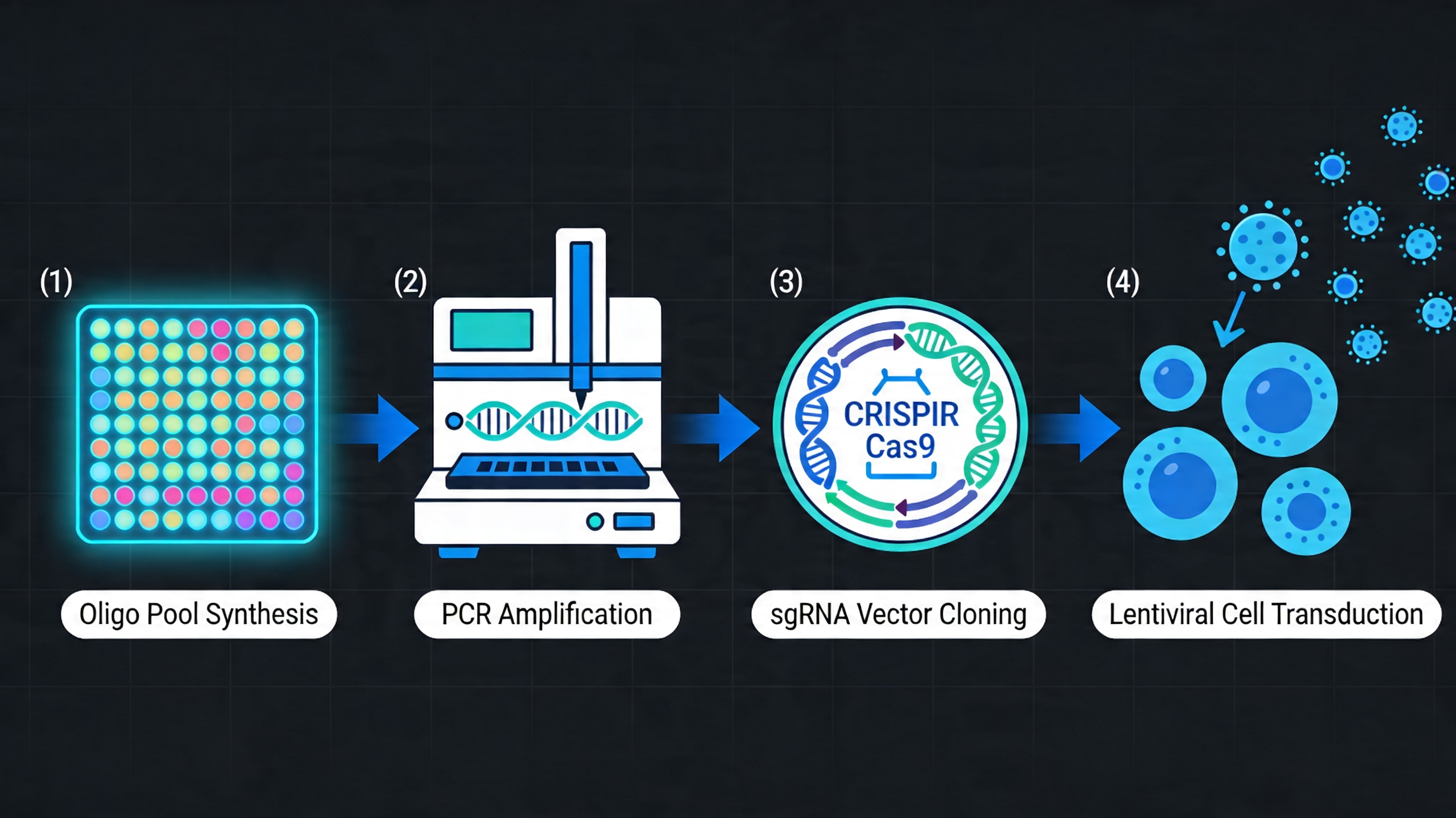
The most established large-scale application of oligo pools is the synthesis of pooled CRISPR guide RNA libraries. A typical genome-wide library contains 60,000 to 100,000 guide sequences, each 20 nt in length, embedded within a longer oligo that includes PCR primer sites and cloning overhangs compatible with the chosen delivery vector.
Oligo pool synthesis makes this feasible at a cost per guide that would be impossible to achieve through any other synthesis approach. The library is amplified from the pool by PCR, cloned into an sgRNA expression vector, and packaged into lentivirus for pooled cell transduction.
Critically, the quality of the oligo pool at the synthesis stage determines the quality of the library at every subsequent step. Representation skew introduced during synthesis propagates through PCR amplification, cloning, viral production, and cell transduction — compounding at each stage. Starting with a high-uniformity pool is the single most effective intervention for reducing technical noise in a CRISPR screen.
Dynegene's CRISPR sgRNA Library service is built on this same synthesis platform, supporting both pooled guide RNA library synthesis and cloned sgRNA library construction for genome-wide and focused pathway screens.
Variant Library Generation for Protein Engineering
Oligo pools are the synthesis backbone of variant library programs in protein engineering, directed evolution, and antibody optimization. A saturation mutagenesis library for a 100-residue protein domain requires approximately 1,900 unique single-mutant sequences — each oligo encoding a different amino acid substitution at a specific position. A combinatorial library covering multiple mutation sites can require tens of thousands to millions of designed sequences.
Before chip-based oligo pool synthesis was available at accessible cost, this scale of design-controlled mutagenesis was not practical. Degenerate codon synthesis (using NNK or NNS degenerate codons at targeted positions) remained common, but this approach generates all 20 amino acids plus stop codons with unpredictable representation, and cannot produce specific combinations of mutations at multiple positions.
Oligo pool synthesis replaces stochastic mutagenesis with sequence-defined variant design. Every library member is intentional. Combined with deep sequencing readout, this enables comprehensive fitness landscape mapping rather than sampling a random subset of possible mutations.
Dynegene's Variant Library for Protein and Nucleic Acid Engineering service extends this capability to include antibody CDR libraries, codon optimization libraries, and combinatorial mutagenesis programs for enzyme and therapeutic protein development.
Custom NGS Target Enrichment Probe Development
Targeted next-generation sequencing depends on hybridization capture probes that selectively enrich genomic regions of interest from a fragmented DNA library. These probes — also called baits — are biotinylated oligonucleotides that hybridize to complementary target sequences, which are then pulled down using streptavidin magnetic beads before sequencing.
The probes themselves are synthesized oligonucleotides of defined sequence, typically 80 to 120 nt in length, tiled across the target regions. For a panel covering 500 genes with typical exon tiling density, the probe set may contain 50,000 to 200,000 unique sequences. This complexity is directly enabled by oligo pool synthesis.
Oligo pools therefore sit upstream of custom NGS panel production. The designed probe sequences are synthesized as a pool, amplified, and processed through a biotinylation step to produce the final capture probe set. The uniformity of the starting oligo pool directly influences the coverage uniformity of the final sequencing data.
Dynegene offers NGS custom probes synthesized on this same platform, combining AI-driven probe design with a hybrid dsDNA and ssDNA probe strategy to achieve on-target rates and GC bias performance that pure ssDNA probe sets typically cannot match. For teams building targeted sequencing panels, this integration between oligo pool synthesis and probe production provides a vertically consistent workflow from designed sequence to sequencing-ready capture reagent.
For whole exome applications, Dynegene's QuarXeq Whole Exome Sequencing Probes cover over 99% of CCDS, RefSeq, and GENCODE-annotated genes, with on-target rates of 98% or above and industry-leading GC bias control.
MPRA and Regulatory Element Screening
Massively parallel reporter assays (MPRA) are a powerful approach for characterizing the functional activity of regulatory DNA elements at genome scale. In a typical MPRA design, thousands of candidate regulatory sequences — promoters, enhancers, CTCF binding sites, or SNP-containing variants — are each linked to a unique DNA barcode and cloned upstream of a reporter gene. The relative activity of each regulatory element is then quantified by sequencing the barcode population from the reporter mRNA.
The oligo pool provides the synthesis layer for both the regulatory element sequences and their associated barcodes. For a study testing 10,000 candidate regulatory elements with 10 barcodes each, the pool must contain 100,000 designed sequences — exactly the scale that chip-based synthesis enables economically.
This application places high demands on synthesis accuracy, because a single-nucleotide error in a candidate regulatory element produces a sequence with a different functional identity than the intended design. Errors that create or destroy transcription factor binding motifs will generate artifactual functional signal that cannot be distinguished from true biology without independent validation.
FISH and Spatial Transcriptomics
Fluorescence in situ hybridization (FISH) probes for single-molecule FISH (smFISH) and related multiplexed spatial transcriptomics workflows are typically designed as sets of 20 to 50 short oligonucleotides (18 to 22 nt) tiling the mRNA target, each conjugated to a fluorophore or a secondary detection handle. For multiplexed spatial assays targeting dozens to hundreds of genes simultaneously, the probe set complexity can reach tens of thousands of sequences.
Oligo pools make this multiplexing economically feasible. Dynegene's Nucleic Acid Probes and Oligo Synthesis Services support probe design and synthesis for these applications, including standard FISH probes and probes for spatially resolved transcriptomics workflows.
Dynegene has also published work on oligo-FISH for chromosome painting applications — a related approach where oligo pools provide the probe complexity needed to generate chromosome-specific hybridization signals without the biological clones or PCR-amplified products that older chromosome painting methods required.
Synthetic Biology and DNA Data Storage
Large-scale synthetic biology programs — genome-scale gene synthesis, metabolic pathway construction, genetic circuit design — require assembling many designed DNA fragments. Oligo pools provide the synthesis precursors for these assembly workflows, generating the short overlapping oligos that are ligated or assembled into longer constructs through methods such as ligase cycling reaction (LCR) or PCA (polymerase cycling assembly).
Large-scale de novo DNA synthesis at Dynegene leverages the same high-throughput microarray platform that underlies the oligo pool service, enabling production of up to 1 Gb of synthesized DNA per run — a throughput level relevant to genome-scale construction projects.
DNA data storage is an emerging application that uses sequence-defined oligonucleotides to encode digital information in DNA molecules. The density of DNA as a storage medium requires large numbers of distinct sequence-defined constructs, and oligo pool synthesis is the only current technology that can produce this complexity at cost. Dynegene has published in this area; a peer-reviewed study on DNA storage image reconstruction using the platform's synthesis output is available in the Citation and References section of the website.
Key Quality Factors in Oligo Pool Synthesis
Not all oligo pools are equal. When evaluating a synthesis platform, quality should be assessed across five distinct dimensions — each of which has direct consequences for experimental outcomes.
Sequence Accuracy
Sequence accuracy refers to the per-base fidelity of synthesized oligos relative to their designed sequences. Errors introduced during synthesis — substitutions, insertions, and deletions — create library members that do not encode the intended sequence.
The consequences scale with library complexity. In a CRISPR library of 80,000 guides, even a 1-in-500 error rate per oligo means that a meaningful fraction of library members carry corrupted sequences. These corrupted sequences occupy library slots without contributing useful experimental data. In functional screens, they add noise to the statistical analysis of screen hits.
For applications like MPRA where single-base differences alter the functional identity of regulatory elements, the requirement for synthesis accuracy is even more stringent.
Representation Uniformity
Uniformity describes how evenly all designed sequences are present in the synthesized pool. A pool where some sequences are present at 1,000-fold the median and others are present at 0.001-fold the median is effectively a pool with far lower complexity than its design specification suggests.
In practice, representation non-uniformity at the synthesis stage is amplified by every downstream processing step. PCR amplification introduces additional bias favoring sequences with efficient primer binding. Cloning introduces bottlenecks that further compress the low end of the representation distribution. Lentiviral packaging introduces yet another representation filter.
The consequence is that a screen designed to evaluate 80,000 unique guides may in practice only provide useful data for 40,000 to 60,000 of them if synthesis uniformity is poor — and the researcher has no way of knowing which guides were lost until the data is analyzed.
Dynegene explicitly positions accuracy and uniformity as core platform performance specifications, which reflects the operational reality that these two factors determine experimental outcomes more than any other synthesis parameter.
Length Capability
Longer oligo length expands design flexibility. Most standard sgRNA library designs require oligos of 60 to 80 nt. Variant library designs with extended coding regions may require 150 nt or more. FISH probe designs that include detection handles, spacers, and secondary structure considerations may need 80 to 120 nt. Some advanced CRISPR applications — including paired-guide designs, base editing templates, and prime editing pegRNAs — require oligos of 200 nt or longer.
A platform capped at 100 nt cannot serve these applications. Dynegene supports oligos up to 350 nt, which covers the length requirements of virtually all current oligo pool applications, including those with extended construct architectures.
Synthesis Throughput
Maximum synthesis complexity per run determines whether a given library design can be completed in a single production batch. For genome-wide CRISPR screens, this means supporting 60,000 to 120,000 unique sequences per chip. For large combinatorial variant programs or DNA data storage applications, complexity requirements can extend to millions of unique sequences.
Dynegene's platform supports up to 4.35 million unique oligos per chip, generating up to 1 Gb of synthesized DNA per run. This capacity is sufficient for the most demanding library designs currently in use in research and industrial genomics programs.
Turnaround Time
Turnaround time affects experimental timelines directly. A 6-week synthesis wait for a CRISPR library delays every subsequent step — viral production, cell line validation, screen execution, and data analysis — by the same margin.
Dynegene's standard turnaround specifications are: 1 week for oligos up to 150 nt, 2 weeks for 151 to 230 nt, and 3 weeks for 231 to 350 nt. Priority processing options are available for smaller pools. For a research program where the library synthesis is on the critical path, this is a meaningful operational parameter.
How to Design a Better Oligo Pool
A successful oligo pool project begins with design decisions made before synthesis. These decisions determine whether the pool will perform as intended in the downstream workflow.
Consider the following before finalizing the design file:
- Downstream workflow: Is the pool going into PCR amplification and cloning, direct hybridization, gene assembly, or a cell-free system? Each workflow has specific requirements for flanking sequences, adapter compatibility, and length constraints.
- Total oligo length: Calculate the functional sequence length plus all overhead elements — universal primers, restriction sites, barcodes, homology arms, scaffold sequences. Confirm that the total fits within the platform's supported length range.
- Sequence diversity requirements: Define minimum required complexity. For a CRISPR screen, this means the number of guides per gene. For a variant library, this means the coverage of mutation space. For a probe set, this means tiling density and target breadth.
- GC content distribution: Sequences with GC content below 25% or above 75% may synthesize poorly or amplify inconsistently. Flag these in the design file and consider alternative sequence positions or probe chemistry.
- Repetitive elements: Oligos designed against repetitive sequences will produce pools where those sequences are present at elevated copy number due to non-specific hybridization during synthesis. This affects both synthesis quality and downstream application performance.
- Pool format: Should the final pool remain unpooled for custom sub-pool applications, or should it be delivered as a single combined pool? For projects that require sub-pool analysis or multi-step assembly, delivery format matters.
The Technical Resources and Instructions section of the Dynegene website includes design guidance for oligo pool projects across different application types.
How Oligo Pools Connect to Adjacent Workflows
Oligo pools rarely operate in isolation. They are typically the synthesis layer that feeds into one or more downstream workflows, each of which has its own quality requirements and processing steps.
Understanding these connections helps research teams design their oligo pool orders with the full workflow in mind:
- CRISPR screens: Oligo pool → PCR amplification → restriction cloning into sgRNA vector → electroporation → plasmid library QC → lentiviral packaging → cell transduction → selection → deep sequencing. Each step has representation requirements that cascade from pool quality. See Dynegene's CRISPR sgRNA Library service for cloned library options.
- Custom NGS panels: Oligo pool → PCR amplification → biotinylation → hybridization capture validation → panel QC. Probe uniformity from the synthesis pool directly translates into on-target coverage uniformity. Dynegene's NGS custom probe service handles the downstream processing from the synthesized pool to the final biotinylated probe set.
- Gene fragment assembly: Oligo pool → overlap extension PCR or ligase cycling reaction → assembly into full-length constructs → sequence verification. This workflow benefits from the Gene Fragments service for projects where longer assembled constructs are required.
- Variant library programs: Oligo pool → amplification → cloning into display or expression vector → phenotypic screening → deep sequencing. Synthesis uniformity is the primary determinant of variant detection completeness. See the Variant Library service for downstream library construction support.
- Antibody library construction: Oligo pool → assembly into CDR-diversified antibody sequences → display selection → sequencing. Dynegene's Synthetic Antibody Library service provides a complete workflow from oligo pool synthesis through antibody library generation.
This is why oligo pools function as an enabling layer across multiple genomics and synthetic biology pipelines rather than a standalone product. The synthesis quality of the pool sets a ceiling on what every subsequent step can achieve.
When to Use Oligo Pools Instead of Gene Fragments
Oligo pools and gene fragments address different but frequently complementary experimental needs. The choice between them is a design decision, not a preference.
Use oligo pools when:
- The project requires very high sequence diversity (hundreds to millions of unique sequences)
- Sequences are short enough to fit within the synthesis platform's maximum length without assembly
- The downstream workflow begins with amplification or hybridization rather than direct cloning of individual constructs
- Budget per sequence is a primary constraint
Use gene fragments when:
- The project requires a longer, defined double-stranded construct for direct cloning, HDR template use, or single-construct validation
- The target sequence is longer than can be accurately synthesized as a single oligo
- The downstream workflow requires a sequence-verified individual construct rather than a pooled mixture
For projects that begin with broad sequence exploration via an oligo pool and later narrow to a small number of validated hits, the natural progression is to follow up with gene fragments or full gene synthesis for the selected candidates. These two synthesis formats are complementary stages of the same research process.
Selecting an Oligo Pool Synthesis Provider
An effective provider should be evaluated across the full range of factors that determine experimental success — not just cost per oligo. Key considerations include:
- Maximum supported oligo length and accuracy at that length
- Maximum library complexity per run
- Representation uniformity specifications and how they are measured
- Turnaround time by pool size and oligo length
- Quality documentation provided with each order
- Compatibility with downstream applications
- Availability of technical support across the library construction workflow, not just at the synthesis step
Dynegene's FAQ page addresses common questions about platform specifications, order requirements, and application compatibility. The patent and intellectual property page documents the proprietary technology underlying the synthesis platform. For teams with complex or novel project requirements, direct technical consultation is available through the contact page.
Dynegene Oligo Pool Platform: Verified Specifications
Dynegene's oligo pool synthesis platform is built on a next-generation ultra-high-throughput DNA microarray synthesis system with complete proprietary intellectual property. The following specifications are derived from the product page and publicly available technical materials:
|
Parameter
|
Specification
|
|
Maximum oligo length
|
350 nt
|
|
Maximum unique oligos per chip
|
4,350,000
|
|
Synthesis output per run
|
Up to 1 Gb
|
|
Standard turnaround (≤150 nt)
|
1 week
|
|
Standard turnaround (151–230 nt)
|
2 weeks
|
|
Standard turnaround (231–350 nt)
|
3 weeks
|
|
Core quality specifications
|
High accuracy; high uniformity
|
These specifications position the platform for compatibility with the most demanding current oligo pool applications — from genome-wide CRISPR screens and deep mutational scanning programs to large-scale synthetic biology constructs and DNA data storage research.
For teams ready to begin an oligo pool project, the Oligo Pools product page provides ordering information, application notes, and direct access to the technical team.
 NGSHybridization Capture Probe NGS custom probes MRD custom probes QuarStar Human MethylCap Panel QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 QuarStar Pan-Cancer Panel 1000-Gene QuarStar HLA Panel QuarXeq HRD panel QuarXeq Mitochondrial Probes Whole Exome Sequencing Probes QuarStar Human All Exon Probes 4.0 (Heredity) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 1.0 (RNA) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Dynegene Adapter Family Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus3 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Blocker Family QuarHyb DNA Reagent Kit Pro(Methyl) Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System
NGSHybridization Capture Probe NGS custom probes MRD custom probes QuarStar Human MethylCap Panel QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 QuarStar Pan-Cancer Panel 1000-Gene QuarStar HLA Panel QuarXeq HRD panel QuarXeq Mitochondrial Probes Whole Exome Sequencing Probes QuarStar Human All Exon Probes 4.0 (Heredity) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 1.0 (RNA) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Dynegene Adapter Family Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus3 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Blocker Family QuarHyb DNA Reagent Kit Pro(Methyl) Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System Primers and Probes
Primers and Probes RNA SynthesissgRNA miRNA siRNA
RNA SynthesissgRNA miRNA siRNA



 Gene
Gene Oligo Pools
Oligo Pools CRISPR sgRNA Library
CRISPR sgRNA Library Antibody Library
Antibody Library Variant Library
Variant Library





 Tel: 400-017-9077
Tel: 400-017-9077 Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai
Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai Email:
Email: Tel: 400-017-9077
Tel: 400-017-9077 Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai
Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai Email:
Email: 







