Hybridization capture is the target enrichment strategy underlying most clinical-grade and research-grade targeted next-generation sequencing panels in use today. Its operational logic is straightforward: biotinylated oligonucleotide probes hybridize to complementary sequences in a fragmented DNA library, and the resulting probe-target duplexes are physically separated from the rest of the library using streptavidin magnetic beads. The captured fraction is then amplified and sequenced at high depth.
What is less widely understood is that every custom capture probe set begins as a designed list of oligonucleotide sequences — and that those sequences are produced using precisely the same chip-based synthesis technology as oligo pools. The path from a BED file of target coordinates to a validated, biotinylated, sequencing-ready probe set runs directly through the oligo pool synthesis step. Understanding this connection is essential for research teams making decisions about custom panel development — who to work with, what specifications matter, and where the technical quality levers are in the production process.
How Oligo Pool Synthesis and Capture Probe Production Are Connected
A hybridization capture probe set and a CRISPR sgRNA library derived from an oligo pool share the same synthesis origin: a DNA microarray platform that synthesizes thousands to millions of sequence-defined oligonucleotides in parallel. Both begin as designed sequences submitted to a chip-based synthesis run. Both are delivered initially as single-stranded oligonucleotides representing all designed sequences in the pool.
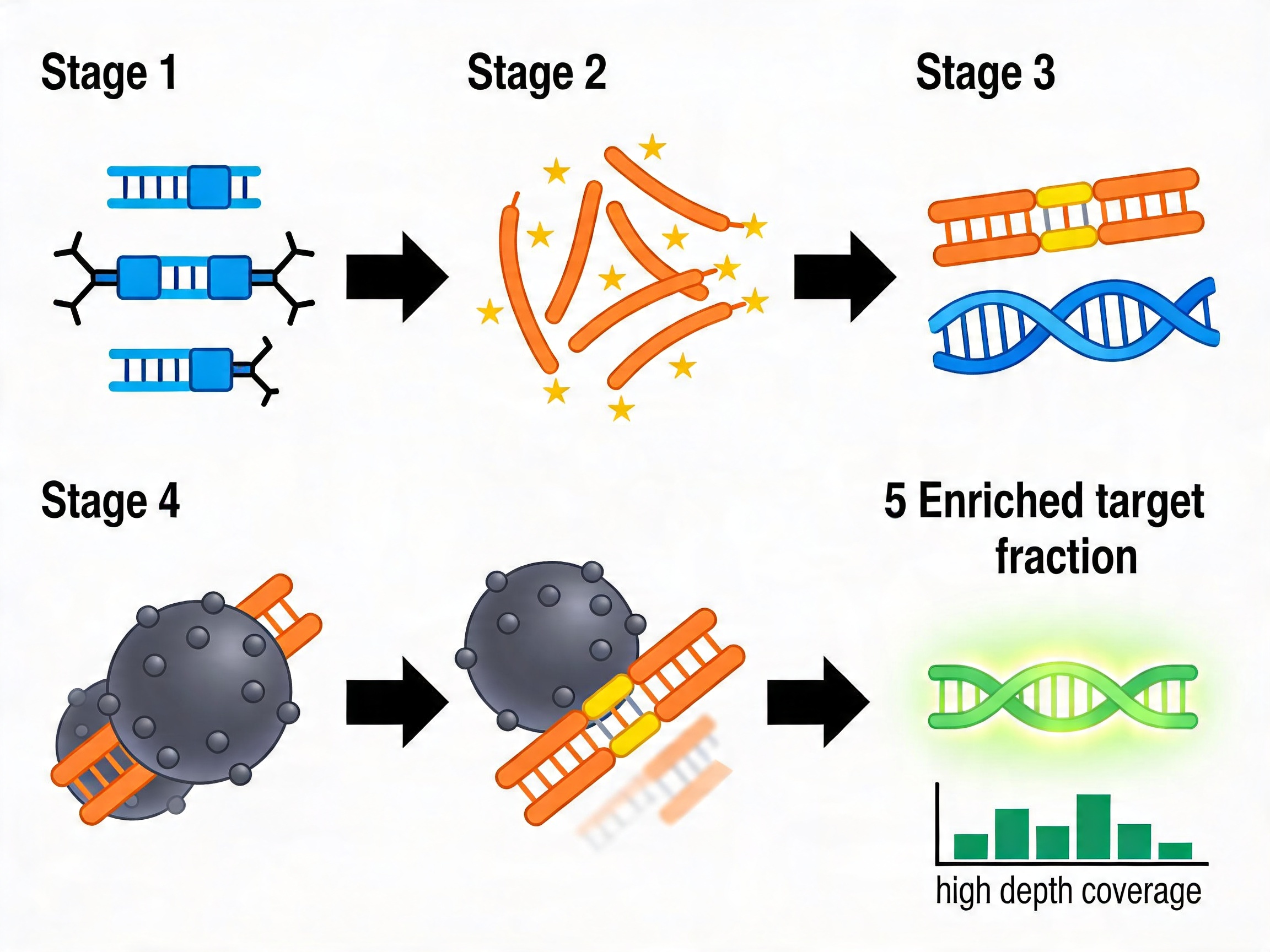
The divergence occurs at the post-synthesis processing step. A CRISPR library pool is amplified, cloned into an expression vector, and packaged into virus. A capture probe set is amplified, biotinylated, and quality-controlled for probe size distribution and labeling efficiency. The downstream processing differs; the synthesis quality requirements are identical.
This matters because the synthesis parameters that define oligo pool quality — sequence accuracy per base, representation uniformity across the pool, maximum supported oligo length, and synthesis throughput — directly determine the probe set quality that any downstream processing step can achieve. No downstream operation can correct for a synthesis pool with high representation skew. A target region that is poorly represented in the synthesis pool will be effectively under-probed in the final panel, regardless of how well the biotinylation and hybridization steps are executed.
Dynegene's Oligo Pools synthesis platform supports up to 4.35 million unique sequences per chip at lengths up to 350 nt, generating up to 1 Gb of synthesized DNA per run — the synthesis infrastructure that underlies both the standalone oligo pool service and the custom probe products in the NGS portfolio.
Why Oligo Pool-Based Custom Panel Development Outperforms Alternatives
Design Latitude That Commercial Panels Cannot Match
Every commercial NGS panel fixes its target content at the point of product validation. The probe sequences, genomic coordinates, tiling strategy, and coverage boundaries are defined by the vendor and cannot be modified by the end user. For programs where the scientific question maps cleanly onto standard commercial panel content, this is acceptable. For programs that do not, it is a fundamental constraint.
Custom oligo pool-derived panels give the research team complete control over every design parameter: the exact chromosomal coordinates and reference assembly version, probe length, tiling density per region, strand balance, boosting density at problematic loci, and the inclusion of any spike-in content — population-specific variant sites, non-coding regulatory regions, deep intronic splice-altering positions, pathogen-derived sequences, or synthetic calibration targets.
This design latitude is operationally critical for applications including:
- Disease-specific panels covering a curated gene set not present in standard commercial offerings
- Research programs requiring coverage of UTRs, promoters, enhancers, or intronic regions beyond canonical exon boundaries
- Infectious disease and metagenomic applications targeting pathogen genomes for which no commercial panel exists
- Multi-species genomics where the reference genome does not correspond to any available commercial panel
- Programs incorporating population-specific variant hotspots for non-European ancestry cohorts
Cost Structure That Favors Scale
Commercial panel reagents are priced on a per-sample basis. Custom probe synthesis has a higher upfront synthesis and validation cost that is amortized across every sample the probe lot is used on. The break-even point depends on per-sample commercial panel reagent cost and the number of samples in the program, but for programs running hundreds to thousands of samples, custom synthesis is economically favorable.
An additional cost efficiency comes from panel leanness. A custom panel that covers only the gene set relevant to the specific scientific question — rather than a broad commercial panel that covers hundreds of additional genes the study will never analyze — reduces total target region size and the sequencing depth required for complete target coverage. Every base of off-target sequence that a leaner panel eliminates represents sequencing cost returned to useful data.
Iteration Speed Compatible With Research Timelines
When a panel needs to be revised — adding newly characterized disease loci, adjusting tiling density based on pilot sequencing data, or extending coverage to a new gene family — the revision requires a new synthesis run. On Dynegene's platform, standard turnaround for oligos up to 150 nt is 1 week, for 151–230 nt is 2 weeks, and for 231–350 nt is 3 weeks. This timeline is compatible with research project revision cycles in a way that alternative probe production methods based on biological amplification or bead-based synthesis are not.
The Complete Production Workflow: From Designed Sequence to Biotinylated Probe Set
Stage 1: Target Region Definition and Probe Sequence Design
The starting material for all custom probe production is a list of target genomic coordinates. For human panels, this is typically provided as a BED file referencing a specified assembly version (GRCh38/hg38 is current standard; GRCh37/hg19 remains in use in clinical genomics programs). For non-human panels, equivalent coordinate formats against the relevant reference are used.
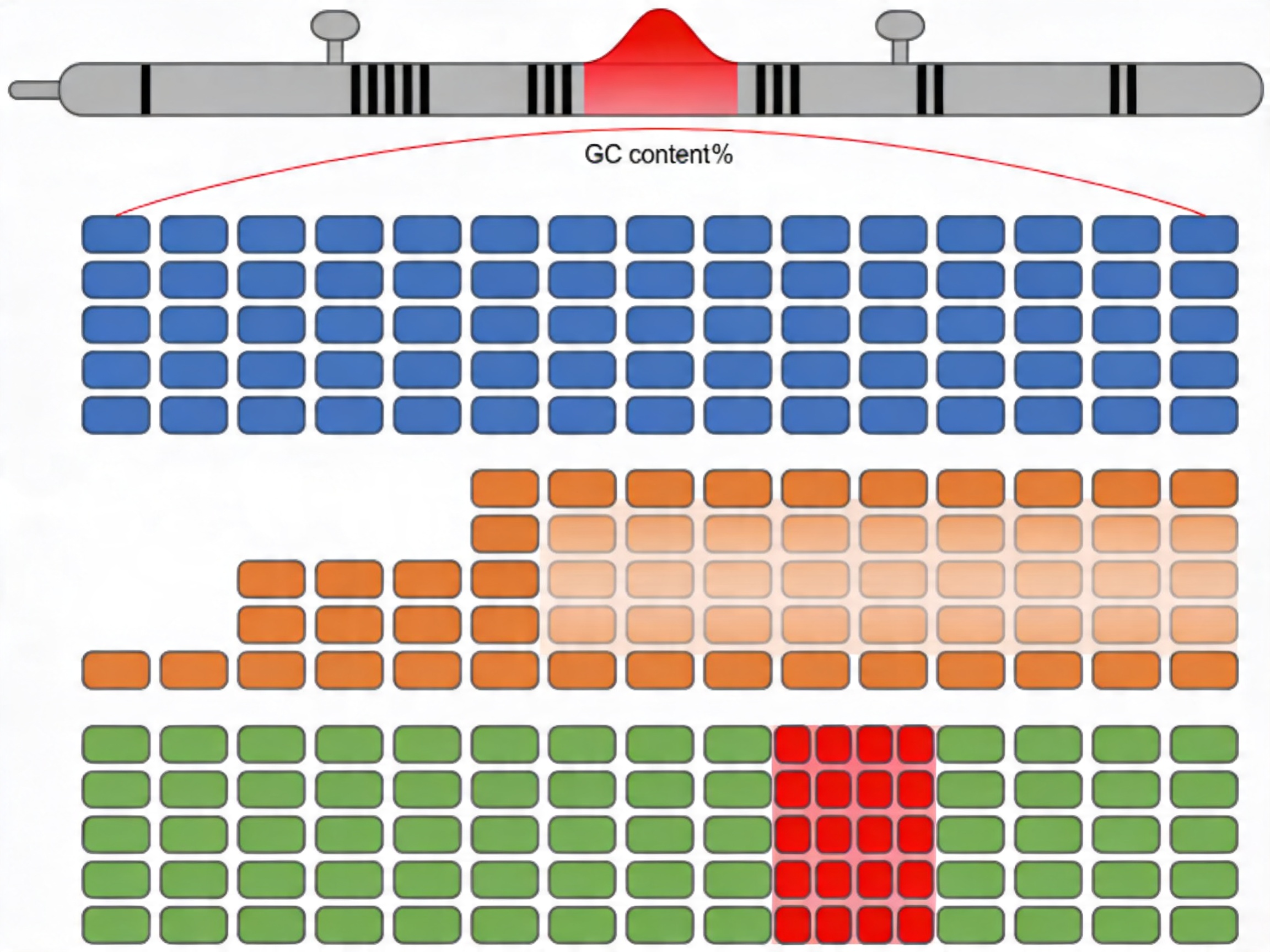
From the target coordinates, probe sequences are designed by tiling the reference sequence at defined spacing. Several design parameters are resolved at this stage and cannot be changed retrospectively without re-synthesis:
Probe length: Most hybridization capture applications use 80 to 120 nt probes. Within this range, longer probes have higher melting temperature (Tm) and more stable probe-target duplexes, which tolerates higher stringency wash conditions and reduces non-specific retention. Shorter probes within this range provide more design positions per kilobase in regions where probe placement is constrained by GC content or repeat proximity.
Tiling strategy: End-to-end (contiguous, no overlap), step tiling with defined overlap (commonly 20–60 nt overlap between adjacent probes), or boosted tiling (standard density supplemented by additional probes at specific difficult positions). The choice affects both total probe count and coverage uniformity per target base. For panels expected to perform well on cfDNA (mononucleosomal fragment length ~160–200 bp), tiling must ensure that even the shortest expected library fragments have sufficient probe overlap length for productive hybridization.
GC content management: Probes with GC content below ~25% or above ~75% have significantly altered Tm relative to the target hybridization temperature. These probes hybridize with reduced efficiency and contribute to coverage dropout in the final data. In silico probe design should flag GC-extreme positions and apply compensatory strategies — increased tiling density, alternate strand probes, or modified probe chemistry — before the design is finalized.
Repeat element masking: RepeatMasker analysis against the relevant reference genome should be applied to all probe sequences before submission. Probes with significant overlap against annotated SINEs, LINEs, simple repeats, or low-complexity sequences will hybridize non-specifically to multiple genomic locations, reducing on-target rates and inflating sequencing cost. Positions with ≥40% repeat overlap typically require redesign.
Pseudogene cross-reactivity: RepeatMasker does not identify pseudogene similarity. A secondary BLAST-based analysis is required for genes with known processed or segmental duplication pseudogenes — common in gene families including globins, SMN, PMS2, and many others relevant to rare disease genomics.
Dynegene's NGS custom probes service applies AI-driven probe design across all of these parameters, using the latest RefSeq, CCDS, and GENCODE annotation databases as the primary target region reference.
Stage 2: Oligo Pool Synthesis
The finalized probe sequence list is submitted to the synthesis platform. The output is a pool of single-stranded oligonucleotides at approximately 2 to 350 nt per sequence, representing every designed probe. At this stage, the pool is a mixture of all probe sequences in a phosphate buffer.
Two key quality metrics are assessed at this stage before proceeding to downstream processing:
Representation uniformity: Quantified by shallow sequencing of the raw pool or by capillary electrophoresis-based methods. Uniformity is typically expressed as the percentage of designed sequences present within a defined fold-window of the median (e.g., percentage of probes within 10-fold of median) or as the skew ratio (90th percentile count / 10th percentile count). A panel with a synthesis-level skew ratio above 5.0 will produce downstream coverage non-uniformity that cannot be corrected by increasing sequencing depth.
Sequence accuracy: Synthesis errors — substitutions, insertions, deletions — generate probe molecules that differ from the designed sequence. For capture probes, a probe with a 2 nt deletion near its center will have a significantly reduced Tm and may fail to hybridize under standard capture conditions. Synthesis error rates compound over oligo length; platforms that maintain accuracy across the full supported length range (up to 350 nt at Dynegene) are necessary for long-probe applications.
Stage 3: PCR Amplification and Double-Stranding
The synthesized pool is amplified by PCR using universal primers corresponding to flanking sequences added to each probe during design. This step converts single-stranded oligos into double-stranded amplicons and increases the total DNA mass available for downstream biotinylation.
Cycle number management at this stage is directly analogous to CRISPR library amplification: each additional cycle provides opportunity for preferential amplification of sequences with favorable primer binding or reduced secondary structure, at the expense of GC-extreme probes. Maintaining amplification below 18–20 total cycles limits the amplification-induced contribution to probe representation skew. High-fidelity proofreading polymerase (Q5, Phusion, or equivalent with error rate ≤1×10/base/duplication) is required to avoid introducing PCR-generated sequence errors into the probe pool.
For probe designs covering GC-rich target regions, betaine (1–2 M final concentration) and DMSO (2–5% v/v) additives in the PCR reaction reduce differential amplification efficiency between AT-rich and GC-rich probe sequences.
Stage 4: Biotinylation — The Mechanistic Choice Between ssDNA and dsDNA Strategies
Biotinylation is the chemical step that enables streptavidin-mediated pulldown. The biotin-streptavidin interaction (Kd ≈ 10⁻¹⁵ M) is among the strongest non-covalent interactions in biochemistry and is stable under the stringent wash conditions used to remove non-specifically hybridized fragments.
Two principal biotinylation strategies are used in commercial probe production, and their choice has performance consequences:
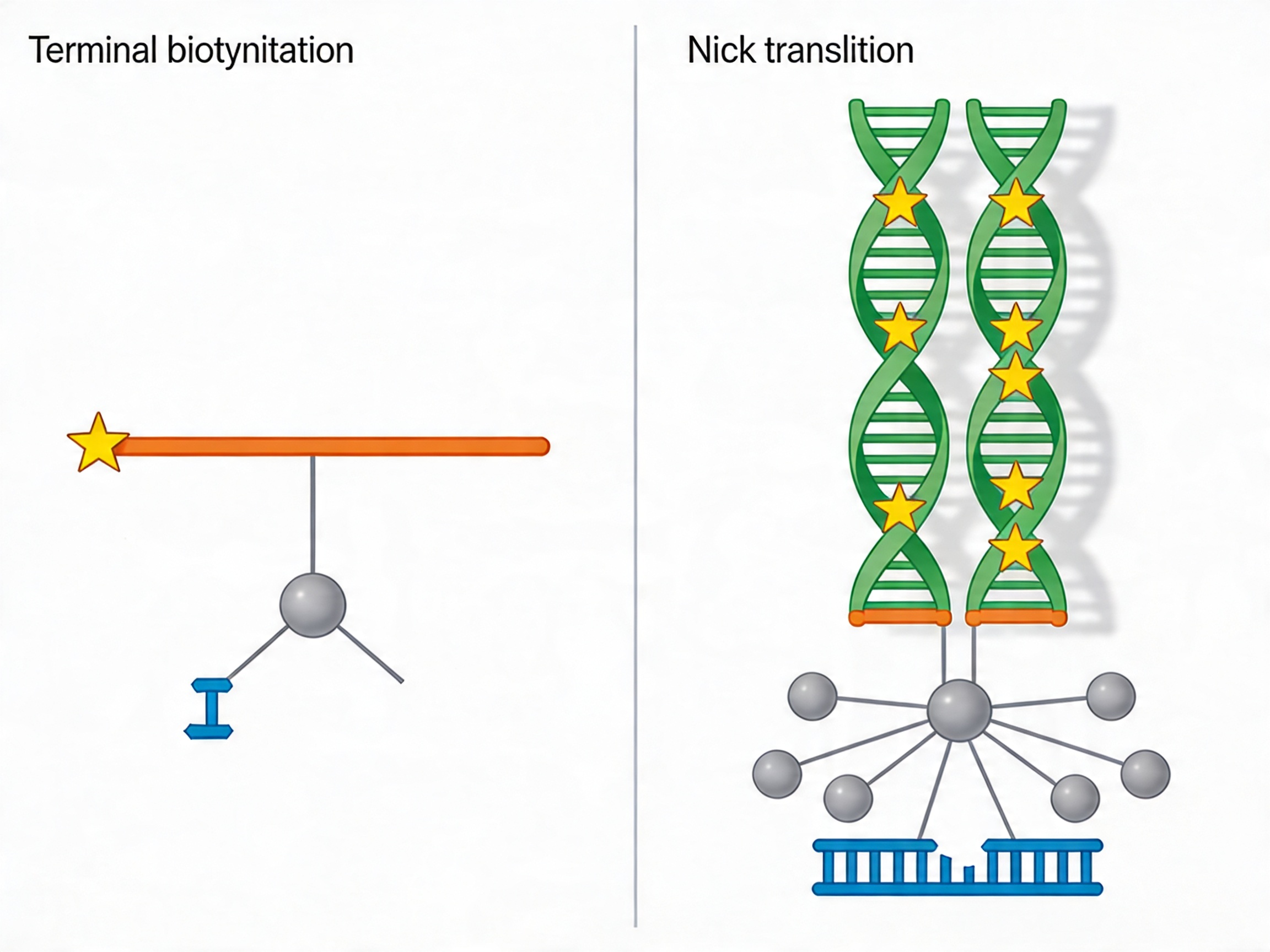
Terminal biotinylation of ssDNA probes: Biotin is incorporated at the 5′ or 3′ terminus during chip-based synthesis using a biotinylated phosphoramidite. This produces one biotin molecule per probe. Terminal biotinylation is straightforward, reproducible at high throughput, and is the standard approach for ssDNA probe production. The limitation is single-point attachment: if the biotin-streptavidin bond is sterically hindered during hybridization or wash steps, the entire probe-target duplex is lost.
Nick translation biotinylation of dsDNA probes: Biotin is incorporated throughout the probe body using biotinylated dNTPs during a nick translation reaction on the double-stranded amplified probe pool. This produces multiple biotin incorporation sites per probe molecule — typically 5 to 15 per 100 nt of probe — providing redundant streptavidin attachment and more robust pulldown efficiency. dsDNA probes also capture targets through strand displacement hybridization, a mechanism that is less susceptible to intramolecular secondary structure competition than the complement hybridization mechanism of ssDNA probes. This is the technical basis for the demonstrated performance advantage of dsDNA probes at GC-rich target sequences.
For RNA probe-based hybridization capture (used in some cDNA library capture and RNA-seq enrichment workflows), Dynegene's Hybridization Capture RNA Probe products provide RNA:DNA duplex hybridization capability with better sensitivity and wider coverage.
Stage 5: Purification and QC Release
Post-biotinylation purification removes unincorporated biotinylated nucleotides (for nick translation products), residual primers, and synthesis byproducts. The purified probe set undergoes QC assessment covering:
- Size distribution confirmation by Bioanalyzer or capillary electrophoresis
- Biotinylation efficiency assessment (HABA assay or functional streptavidin binding test)
- Probe concentration determination by fluorometric quantification
- Optional: shallow sequencing of the biotinylated pool to confirm representation uniformity is maintained through the production process
The probe set is released only on passing all QC criteria. For custom panels, release QC data is provided to the customer as part of the delivery documentation.
Integration with the Complete NGS Library Preparation and Capture Workflow
The probe set is one component of a validated workflow. Its performance in sequencing depends on compatibility with every upstream and downstream step.
Library preparation: The fragmented DNA library must be adapter-ligated and compatible with the blocking reagents and hybridization buffer used in the capture step. Dynegene's DNA Library Preparation Kit — validated for gDNA, FFPE-derived DNA, and cfDNA — produces libraries compatible with the Dynegene probe portfolio and capture reagent system. For enzymatic fragmentation, the Fragmentation Reagent provides controlled fragment size targeting.
Adapter blocking: During hybridization, adapter sequences on library fragments can hybridize to each other, creating adapter-dimers that compete for streptavidin binding and reduce on-target capture efficiency. Blocking oligonucleotides complementary to the adapter sequences are included in the hybridization reaction to prevent this. Dynegene's Blocker Family provides blocking reagents validated for use with Dynegene adapter chemistries.
Hybridization buffer and conditions: The hybridization temperature, salt concentration, and formamide content of the hybridization buffer determine the stringency of probe-target duplex formation and influence both on-target rate and GC bias. Dynegene's QuarHyb Super DNA Reagent Kit and QuarHyb DNA Plus 3 Reagent Kit provide optimized hybridization formulations validated for use with the Dynegene probe portfolio.
Streptavidin pulldown: Streptavidin magnetic beads with validated binding capacity and wash performance are required for consistent capture efficiency. Bead quality — specifically the consistency of streptavidin surface density and non-specific DNA binding levels — affects both on-target rate and duplicate rate in the final sequencing data.
Application-Specific Panel Design Considerations
Oncology Somatic Variant Panels
Somatic variant detection at low allele fractions — circulating tumor DNA at 0.1–1.0% VAF, intratumor heterogeneity subclones — requires on-target rates ≥90% and mean target depth ≥500× for reliable variant calling. Custom panel design for these applications prioritizes high probe uniformity and low off-target background over breadth of target coverage. Panels should include MSI marker loci and structural variant breakpoint regions in addition to point mutation hotspots. Dynegene's QuarStar Pan-Cancer series, QuarStar Liquid Pan-Cancer Panel 3.0, and QuarStar Pan-Cancer Fusion Panel 1.0 represent validated commercial configurations for these applications. Custom spike-in probes can extend any of these panels with additional targets.
Rare Disease and Mendelian Genetics Panels
Gene panels for rare disease diagnosis require complete coverage of canonical coding exons, canonical splice donor and acceptor sites (typically ±2 nt of exon boundaries), and clinically characterized intronic variants. For genes with pseudogene paralogs — including PMS2, SMN1/2, CYP2D6, and others — targeted design approaches are required to maintain locus specificity. Custom panels allow the gene content to be matched precisely to the diagnostic application and updated as new disease genes are identified.
Whole Exome Sequencing
For programs requiring comprehensive exome coverage as a baseline, Dynegene's Whole Exome Sequencing Probes provides approximately 99.5% coverage of CCDS, RefSeq, and GENCODE core coding regions. The QuarStar Human All Exon Probes 4.0 series — available in Heredity, Tumor, and Standard configurations from the custom probe service — extends WES capability with application-specific content optimization. Custom spike-in augmentation adds program-specific targets without replacing the validated exome backbone.
Pathogen and Metagenomic Capture
Hybridization capture for pathogen enrichment from clinical samples uses probes designed against pathogen reference genomes rather than human genomic coordinates. This application benefits from the same design principles — tiling strategy, GC management, cross-reactivity avoidance — but requires reference sequences from pathogen genome databases (NCBI RefSeq viral/bacterial genomes, PATRIC, etc.) rather than GRCh38. Commercial pathogen panels such as Dynegene's PathoSeq 450 Pathogen Library cover a broad spectrum of clinically relevant pathogens; custom probe design extends this to novel or outbreak-specific organisms.
Methylation Capture Panels
For targeted bisulfite sequencing or methylation-sensitive capture applications, probe design must account for the bisulfite conversion of unmethylated cytosines to uracil (read as thymine). Probes designed against unconverted reference sequences will hybridize with reduced efficiency to bisulfite-converted libraries due to the C→T changes at unmethylated positions. Bisulfite-compatible probe design uses converted reference sequences and requires strand-specific design to maintain specificity. Dynegene's QuarStar Human MethylCap Panel is specifically designed for ultra-large human methylome research.
Realistic Development Timeline for Custom Panel Projects
| Stage |
Typical Duration |
| Probe design, repeat masking, GC analysis, and design file finalization |
2–3 days |
| Probe synthesis (≤150 nt) |
2–3 weeks |
| QC |
7–10 days |
Total elapsed time from project initiation to a validated, production-ready custom panel is typically 5 to 6 weeks for focused panels and 6 to 7 weeks for complex multi-application panels requiring multiple design-test cycles.
What to Provide When Initiating a Custom Panel Project
To enable accurate design proposal, quote, and production timeline, provide the following at project initiation:
- Target region coordinates in BED format, with reference assembly version specified
- List of any known problematic regions (high-GC gene families, pseudogene-containing loci, known repeat-adjacent targets)
- Sample type and DNA input quality (high-quality gDNA, FFPE, cfDNA, low-input)
- Required minimum on-target rate and coverage uniformity specifications
- Sequencing platform and expected read length
- Whether spike-in augmentation of a standard Dynegene panel is preferred, or a fully custom design is required
- Expected sample volume over the program lifetime (for cost structure assessment)
For complex or non-standard panel requirements, technical consultation before project submission improves design quality and reduces revision cycles. Dynegene's team is available through the Contact page.
Published performance benchmarks for Dynegene's NGS probe products — including comparative WES performance data from Shanghai Jiaotong University and validation data for the pan-cancer panel series — are available in the Citation and References section. Platform specifications, ordering formats, and technical FAQs for oligo pool and probe synthesis projects are documented at Technical Resources.
 NGSHybridization Capture Probe NGS custom probes MRD custom probes QuarStar Human MethylCap Panel QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 QuarStar Pan-Cancer Panel 1000-Gene QuarStar HLA Panel QuarXeq HRD panel QuarXeq Mitochondrial Probes Whole Exome Sequencing Probes QuarStar Human All Exon Probes 4.0 (Heredity) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 1.0 (RNA) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Dynegene Adapter Family Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus3 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Blocker Family QuarHyb DNA Reagent Kit Pro(Methyl) Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System
NGSHybridization Capture Probe NGS custom probes MRD custom probes QuarStar Human MethylCap Panel QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 QuarStar Pan-Cancer Panel 1000-Gene QuarStar HLA Panel QuarXeq HRD panel QuarXeq Mitochondrial Probes Whole Exome Sequencing Probes QuarStar Human All Exon Probes 4.0 (Heredity) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 1.0 (RNA) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Dynegene Adapter Family Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus3 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Blocker Family QuarHyb DNA Reagent Kit Pro(Methyl) Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System Primers and Probes
Primers and Probes RNA SynthesissgRNA miRNA siRNA
RNA SynthesissgRNA miRNA siRNA



 Gene
Gene Oligo Pools
Oligo Pools CRISPR sgRNA Library
CRISPR sgRNA Library Antibody Library
Antibody Library Variant Library
Variant Library





 Tel: 400-017-9077
Tel: 400-017-9077 Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai
Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai Email:
Email: Tel: 400-017-9077
Tel: 400-017-9077 Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai
Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai Email:
Email: 







