Gene fragments and full-length gene synthesis address overlapping but distinct experimental needs. Treating them as interchangeable — either always defaulting to full gene synthesis for convenience, or always using fragments to save cost — leads to avoidable workflow problems: unnecessary expense, sequence gaps that prevent direct cloning, or constructs that cannot be used in the intended downstream assay without additional processing.
This guide provides a technically grounded framework for choosing between the two formats, covering the biological rationale, length constraints, assembly compatibility, and the scenarios where each approach is the appropriate choice. It also explains how gene fragments and oligo pools work together within the same research program, and when neither format alone is sufficient.
What Are Gene Fragments
A gene fragment is a double-stranded, sequence-defined DNA construct that represents a portion of a larger gene, regulatory element, or functional construct — not a complete open reading frame. The distinguishing characteristic is that it is synthesized de novo to an exact user-specified sequence, delivered as a purified double-stranded DNA product ready for direct downstream use.
This is what separates gene fragments from both PCR products and oligonucleotides. A PCR product requires an existing template; a gene fragment does not. An oligonucleotide is single-stranded and typically below 200 nt; a gene fragment is double-stranded and can cover several hundred base pairs of sequence, depending on the synthesis platform.
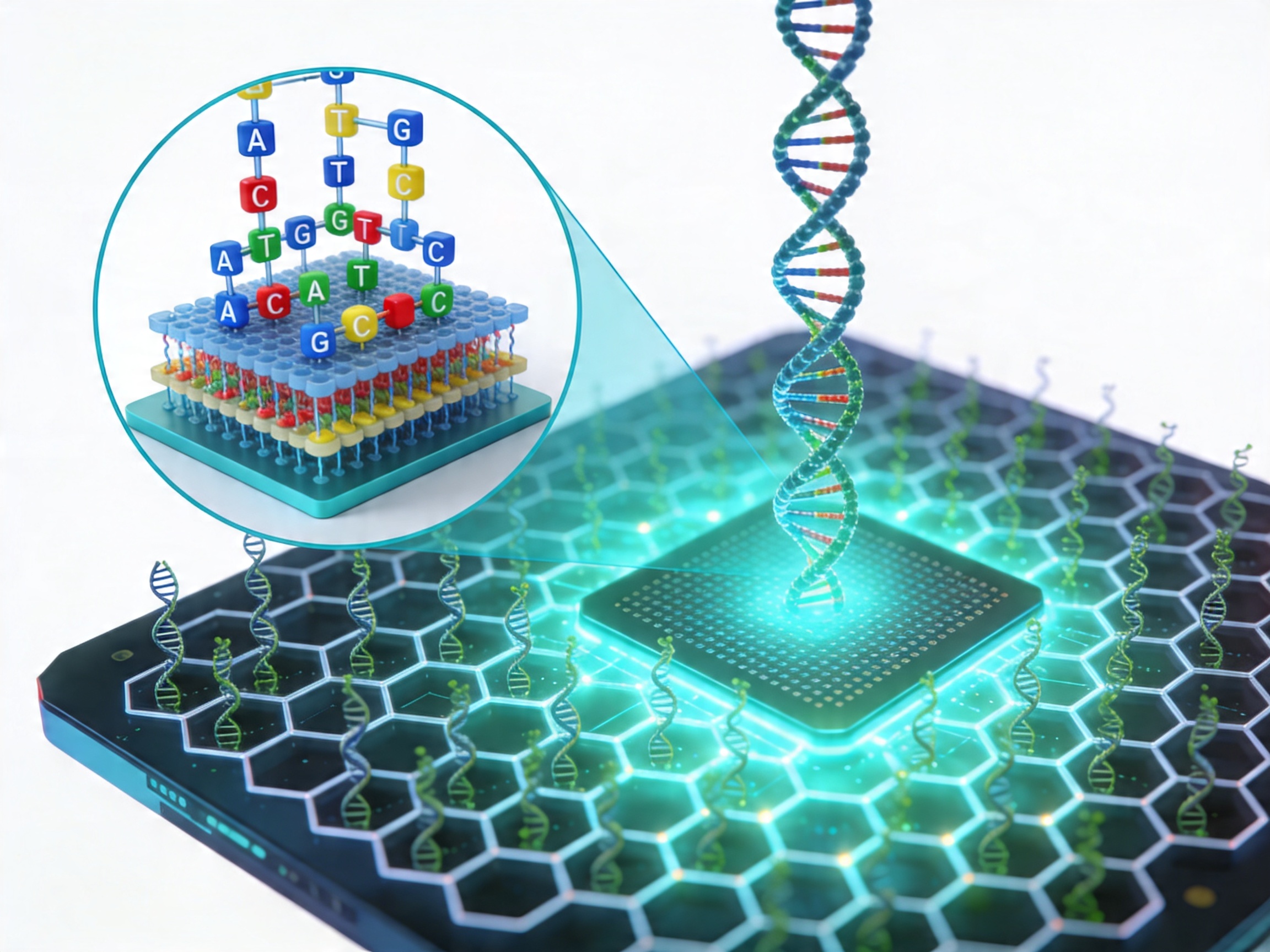
The practical implication is that gene fragments can be used directly in restriction enzyme cloning, Gibson Assembly, Golden Gate Assembly, or as HDR donor templates without a pre-existing template DNA — making them uniquely useful for novel sequence designs, synthetic regulatory regions, codon-modified coding sequences, and sequence variants that do not exist as amplifiable templates in any current plasmid or genomic DNA source.
Dynegene's gene fragment synthesis service is powered by a honeycomb pixel-based ultra-high-throughput DNA microarray synthesis platform, the same platform underlying the oligo pools service. This architecture supports oligos up to 350 nt per position, up to 4.35 million unique oligos per chip, and up to 1 Gb of synthesized DNA per run — enabling both high-throughput parallel fragment production and rapid turnaround for individual projects.
What Is Full-Length Gene Synthesis
Full-length gene synthesis is the de novo production of a complete coding sequence, gene construct, or large defined DNA molecule — typically ranging from a few hundred base pairs to several kilobases. The synthesis process involves designing overlapping oligonucleotides that cover the full-length target sequence, assembling them by polymerase chain assembly or ligase cycling reaction, and then sequence-verifying the assembled product before delivery.
The critical distinction from gene fragments is functional completeness. A full-length synthesized gene is delivered as a single defined construct that encodes everything needed for the intended application — the complete open reading frame, any required regulatory elements that must be present in the final product, and codon optimization for the target expression host if specified.
Dynegene's Gene Synthesis service covers standard gene synthesis as well as Gene Cloning services for constructs that need to be delivered in a specific vector context, verified by sequencing, and ready for direct transformation or transfection.
Comparing the Two Formats
| Dimension |
Gene Fragments |
Full-Length Gene Synthesis |
| Product form |
Double-stranded DNA, partial sequence |
Complete coding sequence or construct |
| Typical length range |
Up to ~600 bp depending on assembly |
Hundreds bp to several kb |
| Template requirement |
None — de novo synthesis |
None — de novo synthesis |
| Codon optimization |
Applicable to the fragment region |
Applicable across the full ORF |
| Sequence verification |
Per fragment |
Delivered as sequence-verified full construct |
| Direct expression |
Not without assembly or vector context |
Yes, with appropriate vector |
| Cost |
Lower per project for partial sequences |
Higher, scales with length and complexity |
| Turnaround |
Faster for shorter constructs |
Longer, especially above 2 kb |
| Primary use cases |
HDR templates, assembly inputs, domain swaps, variant screening |
Expression constructs, novel gene design, pathway engineering |
When Gene Fragments Are the Appropriate Choice
HDR Donor Templates for CRISPR Genome Editing
Gene fragments are among the most commonly requested synthesis products for CRISPR-based precise genome editing. When using homology-directed repair (HDR) to introduce a defined sequence change — a point mutation, an in-frame tag insertion, a regulatory element replacement, or a coding sequence swap — the donor template must present the intended edit flanked by homology arms that match the endogenous sequence flanking the cut site.
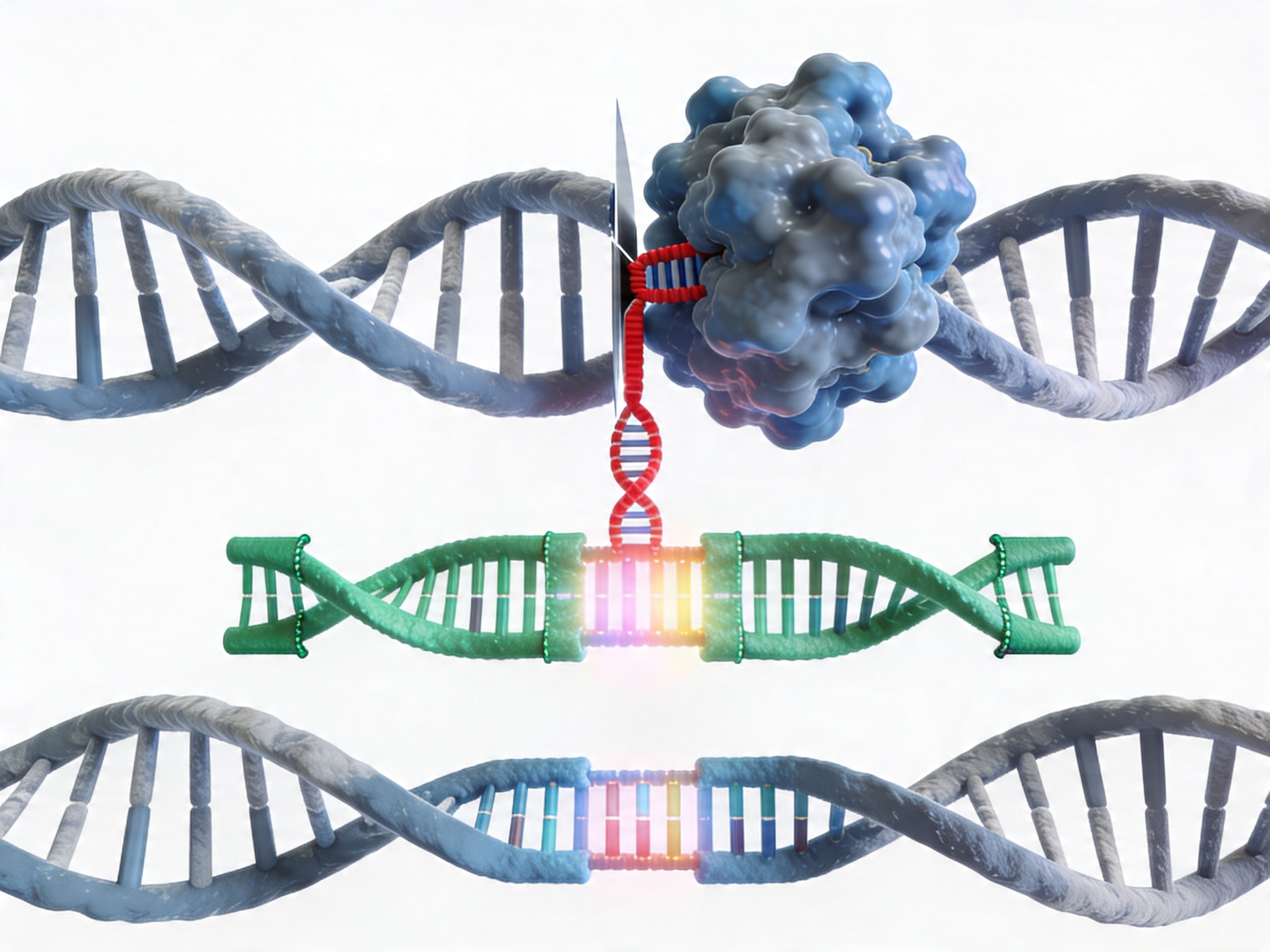
For most HDR applications in cell lines, single-stranded oligonucleotides (ssODNs) are used for small edits within approximately 50 bp of the cut site. When the intended edit is larger — inserting a fluorescent protein tag, replacing an exon, or introducing a synthetic regulatory sequence — a double-stranded DNA donor template of several hundred base pairs is required. A gene fragment synthesized to the exact design specification is the cleanest way to produce this donor without requiring a pre-existing plasmid template.
The homology arms in the donor should ideally be 400 to 800 bp on each side of the edit for plasmid-based donors, or 50 to 150 bp for shorter dsDNA fragments used in linear donor strategies. The specific length and the strand configuration of the donor influence HDR efficiency and the frequency of non-homologous end joining of the donor itself — parameters that should be determined empirically for the specific cell type and locus being targeted.
Dynegene's gene fragment service supports synthesis of these donor constructs to user-specified sequence, with the exact start and end positions, homology arm sequences, and edit-containing internal sequence defined at order submission.
Multi-Fragment Assembly in Synthetic Biology and Metabolic Engineering
In synthetic biology projects, complete gene constructs, biosynthetic pathway elements, or synthetic genetic circuits are frequently assembled from multiple shorter fragments rather than ordered as a single full-length synthesis. The rationale is modularity: individual fragments can be designed, synthesized, and validated independently before being joined in a final assembly step, reducing the risk that a single synthesis failure or design error invalidates the entire project.
Gibson Assembly is the most widely used method for this workflow. It requires overlapping sequences of 15 to 30 bp between adjacent fragments, with overlaps designed to have a melting temperature of approximately 48°C or higher for efficient exonuclease-mediated joining. Golden Gate Assembly uses Type IIS restriction enzymes (most commonly BsaI or BbsI) that cut outside their recognition sequence, generating custom 4-nt overhangs that are unique across the assembly and direct the fragments into the correct order and orientation in a single ligation reaction.
Gene fragments designed for these assembly methods must include the specified overlap or overhang sequences as part of their design. This is a design step that should be completed before synthesis, not after, because the junction sequences affect both the synthesis oligo design and the final assembled product's sequence.
For large synthetic constructs, Dynegene's Gene Cloning service and high-throughput gene synthesis capabilities provide options for producing and assembling larger constructs.
Domain Function Studies and Partial Sequence Validation
Protein structure-function studies frequently focus on specific domains rather than complete genes. Researchers characterizing a kinase domain, an antibody variable region, an enzyme active site loop, or a transcription factor DNA-binding domain typically need only the sequence encoding that region — not the full protein. Ordering a complete gene synthesis to study a 150 amino acid domain that represents 15% of a 1,000 amino acid protein is neither necessary nor cost-efficient.
Gene fragments enable these focused studies directly. The fragment encoding the domain of interest can be synthesized with flanking restriction sites or assembly overhangs compatible with the expression or display vector being used, and cloned directly into the appropriate context.
The same principle applies to regulatory element studies. A researcher characterizing enhancer activity, promoter strength, or the functional effect of disease-associated SNPs in a non-coding region needs only the regulatory element sequence — typically 200 to 600 bp — synthesized with flanking cloning handles for insertion into a reporter vector.
Variant Screening Before Full Construct Commitment
When a research program involves evaluating multiple sequence designs — alternative codon usages, promoter variants, signal peptide sequences, linker designs, or domain boundary positions — it is common to screen several candidate sequences before committing to full construct production.
Gene fragments allow this screening to proceed at lower cost and faster turnaround than full gene synthesis. A set of 5 to 20 fragment variants encoding alternative designs can be synthesized, cloned, expressed in a small-scale format, and evaluated for the relevant property (expression level, solubility, binding, catalytic activity) before the best-performing design is selected for full construct production.
This staged approach is especially practical when working with variant library programs: an oligo pool provides the synthesis layer for broad parallel screening, identified hits are followed up as individual gene fragments for validation, and validated sequences proceed to full gene synthesis or direct cloning for final construct preparation.
Module Insertion into Existing Vector Backbones
Many laboratories maintain a library of expression vectors optimized for specific host organisms, promoter systems, purification tags, or selection markers. When the only change required is inserting a new coding sequence or replacing an existing module — for example, swapping one antibody variable domain for another, replacing a fluorescent protein tag, or inserting a new enzyme variant — the gene fragment representing only the insert is all that is needed.
Ordering a complete gene synthesis including the vector backbone sequence in this scenario adds cost and turnaround time with no experimental benefit. A correctly designed gene fragment with compatible cloning overhangs is sufficient for direct ligation or assembly.
When Full-Length Gene Synthesis Is the Appropriate Choice
Full gene synthesis is the correct format when:
- The complete open reading frame is required for expression: The protein must be encoded in full, with start codon, stop codon, and all intervening sequence, for the construct to function as an expression unit.
- The target sequence is novel with no reference template: For completely synthetic gene designs — codon-optimized sequences for heterologous expression, synthetic regulatory circuits with designed logic, or artificial gene sequences that have no natural counterpart — full gene synthesis provides the complete and sequence-verified product in a single service.
- Codon optimization across the entire coding sequence is required: Codon optimization for expression in a specific host (E. coli, CHO cells, yeast, insect cells) should be applied to the complete ORF to produce a consistent codon usage profile.
- The construct is too long for direct fragment synthesis and assembly is not feasible: Full gene synthesis services include error correction and sequence verification as part of the production workflow.
- A sequence-verified ready-to-use construct in a specific vector is required: Dynegene's Gene Cloning service delivers the synthesized gene already cloned into a specified vector backbone, sequence-verified, and ready for transformation, transfection, or further cloning.
How Gene Fragments and Oligo Pools Work Together
Gene fragments and oligo pools are not competing formats. They serve different stages of the same research program and are most powerful when used in combination.
Oligo pools are the appropriate format when the project requires parallel synthesis of very large numbers of distinct short sequences — for CRISPR library construction, variant library generation, massively parallel reporter assays, or hybridization capture probe production.
Gene fragments are the appropriate format when the project requires a specific, isolated, double-stranded DNA construct of defined sequence — a validated hit from a screen that needs individual characterization, an HDR donor template for a precision edit, or a modular insert for assembly into a larger construct.
A typical progression within a single research program looks like this: an oligo pool is synthesized for broad screening across thousands of designed sequences. Sequencing data from the screen identifies 5 to 20 candidate sequences with the desired properties. Those candidates are ordered as individual gene fragments for confirmatory validation in the intended assay format. One or two validated candidates proceed to full gene synthesis or direct cloning in the expression vector for final construct production.
At each stage, the synthesis format is matched to the scale and purpose of the experiment. Oligo pools provide breadth; gene fragments provide precision; full gene synthesis provides functional completeness.
Practical Design Considerations Before Ordering
Before submitting a gene fragment order, confirm the following design parameters:
- Total length including all functional elements: Calculate the total fragment length including the coding or functional sequence, any flanking homology arms, restriction sites, assembly overhangs, barcodes, or tag sequences.
- Assembly junction sequences: If the fragment will be used in Gibson Assembly, confirm that all junction overlaps are 15 to 30 bp with appropriate GC content and Tm. If Golden Gate Assembly is used, confirm that BsaI or BbsI recognition sequences are not present within the fragment body.
- Restriction site compatibility: Confirm that the enzyme sites in the flanking sequence are not also present internally in the fragment.
- GC content and secondary structure: Fragments with regions of very high GC content (above 70%) or predicted strong secondary structure may synthesize with lower fidelity.
- Sequence verification requirements: For HDR donor templates and expression constructs, Sanger sequencing verification of the delivered fragment is recommended before proceeding to time-sensitive downstream steps.
For detailed platform specifications and ordering guidance, Dynegene's Technical Resources page provides application notes and protocol information for gene fragment projects.
Dynegene Gene Fragment Synthesis Platform
Dynegene's gene fragment synthesis service uses the same honeycomb pixel-based ultra-high-throughput DNA microarray synthesis infrastructure underlying the oligo pools and CRISPR sgRNA library services.
Verified platform specifications:
| Parameter |
Specification |
| Maximum oligo length per chip position |
350 nt |
| Maximum unique sequences per chip |
4,350,000 |
| Synthesis output per run |
Up to 1 Gb |
| Throughput-optimized mode |
Sub-Pool Synthesis |
| Speed-optimized mode |
Mini-Pool Synthesis |
For research teams running programs that require both oligo pools for library-scale synthesis and gene fragments for individual construct validation, this shared platform ensures that both products are produced under the same synthesis quality standards.
Decision Guide
Choose gene fragments when:
- The target sequence is a defined partial region (domain, exon, regulatory element, insert)
- The construct will be used as an HDR template, assembly input, or module swap
- Multiple sequence variants need to be evaluated before committing to full construct production
- The vector backbone and all other elements already exist and only the insert is needed
Choose full gene synthesis when:
- The complete open reading frame with all functional elements is required
- Codon optimization must be applied across the entire coding sequence
- The construct needs to be delivered sequence-verified and cloning-ready in a specific vector
- The target sequence is too long or complex for fragment-based assembly
Consider both in combination when:
- The program begins with broad parallel screening (oligo pool) and proceeds through individual hit validation (gene fragment) to final construct production (full gene synthesis or gene cloning)
For projects where the format choice is unclear based on these criteria, Dynegene's technical team is available through the Contact page to provide guidance specific to the experimental design and downstream workflow.
 NGSHybridization Capture Probe NGS custom probes MRD custom probes QuarStar Human MethylCap Panel QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 QuarStar Pan-Cancer Panel 1000-Gene QuarStar HLA Panel QuarXeq HRD panel QuarXeq Mitochondrial Probes Whole Exome Sequencing Probes QuarStar Human All Exon Probes 4.0 (Heredity) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 1.0 (RNA) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Dynegene Adapter Family Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus3 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Blocker Family QuarHyb DNA Reagent Kit Pro(Methyl) Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System
NGSHybridization Capture Probe NGS custom probes MRD custom probes QuarStar Human MethylCap Panel QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 QuarStar Pan-Cancer Panel 1000-Gene QuarStar HLA Panel QuarXeq HRD panel QuarXeq Mitochondrial Probes Whole Exome Sequencing Probes QuarStar Human All Exon Probes 4.0 (Heredity) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 1.0 (RNA) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Dynegene Adapter Family Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus3 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Blocker Family QuarHyb DNA Reagent Kit Pro(Methyl) Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System Primers and Probes
Primers and Probes RNA SynthesissgRNA miRNA siRNA
RNA SynthesissgRNA miRNA siRNA



 Gene
Gene Oligo Pools
Oligo Pools CRISPR sgRNA Library
CRISPR sgRNA Library Antibody Library
Antibody Library Variant Library
Variant Library





 Tel: 400-017-9077
Tel: 400-017-9077 Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai
Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai Email:
Email: Tel: 400-017-9077
Tel: 400-017-9077 Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai
Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai Email:
Email: 







