Targeted next-generation sequencing concentrates sequencing depth on defined genomic regions, making it the preferred approach for applications where comprehensive, high-depth coverage of a specific gene set matters more than broad but shallow whole-genome data. The capture step that achieves this enrichment is technically straightforward in concept — probes hybridize to target sequences, which are then pulled down magnetically — but the performance of that step is determined almost entirely by decisions made during probe design, before a single reagent is used.
A panel that achieves 98% on-target rate and uniform coverage across all targeted bases does so because the probe set was designed with attention to GC content management, repeat element avoidance, tiling strategy, and probe chemistry. A panel that generates 60% on-target rates and coverage dropout across GC-rich exons fails for the same reasons, applied in reverse.
This guide covers the design principles that distinguish high-performing custom capture panels from those that underperform. It is relevant for research teams building oncology panels, rare disease gene sets, pharmacogenomics panels, whole exome configurations, or any other targeted sequencing application where hybridization capture is the enrichment strategy.
How Hybridization Capture Works: The Physical Chemistry Behind the Assay
Hybridization capture works by exploiting the thermodynamic stability of DNA duplexes. Biotinylated probe oligonucleotides are incubated with a fragmented and adapter-ligated DNA library under conditions that favor stable hybridization between complementary sequences. Probe-target duplexes form preferentially at correctly matched loci. The biotinylated probe-target complexes are then captured using streptavidin-coated magnetic beads — the biotin-streptavidin interaction has one of the strongest non-covalent binding affinities in biochemistry (Kd ≈ 10⁻¹⁵ M) and is resistant to the wash conditions used to remove non-specifically bound fragments.

After capture, the beads are washed under stringent conditions that disrupt weakly hybridized non-specific fragments while maintaining stable on-target duplexes. The captured target sequences are eluted, amplified, and submitted for sequencing.
Several practical points follow directly from this physical picture. First, hybridization stringency — controlled primarily by temperature, salt concentration, and formamide concentration during the hybridization and wash steps — determines the discrimination between on-target and near-target sequences. Second, probe Tm (melting temperature) determines the appropriate hybridization temperature; a probe set with highly variable Tm values across its members will have suboptimal hybridization for some fraction of probes at any single hybridization temperature. Third, the ratio of probe to target in the hybridization reaction influences capture efficiency — excess probe drives the equilibrium toward capture, but very high probe-to-target ratios can also increase non-specific background.
Blocking oligonucleotides are used in the hybridization reaction to prevent adapter sequences from hybridizing to each other or to repetitive sequences that might otherwise capture non-specific library fragments. Dynegene provides a Dynegene Blocker Family of blocking reagents specifically designed for use with its hybridization capture workflows, compatible with the major adapter chemistries used in NGS library preparation.
The downstream capture reagents — including hybridization buffers, wash solutions, and streptavidin magnetic bead systems — are equally important for panel performance. Dynegene's QuarHyb Super DNA Reagent Kit, QuarHyb DNA Plus3 Reagent Kit, and Capture Reagent for DNA Probes Hybridization provide optimized hybridization and wash chemistries validated for use with both DNA capture probes and the specific panel products in the Dynegene portfolio.
Key Parameters in Probe Design
Target Region Definition and Annotation
The first step in probe design is the precise definition of genomic target regions. This requires more than identifying a gene list — it requires specifying exact chromosomal coordinates, strand assignment, and the functional annotation context of each region.
For human genomic panels, target coordinates should be derived from a current, well-curated annotation source. Dynegene's probe design pipeline uses the latest RefSeq, CCDS, and GENCODE databases as the primary annotation references, which together provide near-complete coverage of protein-coding exons and canonical splice sites.
The choice of annotation database matters operationally. RefSeq is the most conservative annotation — it includes only well-supported transcripts. GENCODE is more comprehensive and includes many predicted and lower-confidence transcripts. CCDS defines a high-confidence set of coding sequences with cross-database consensus. Using all three in combination, as Dynegene does, maximizes coverage of functionally characterized regions while avoiding inclusion of unsupported coordinates that may generate probe-wasting non-specific capture.
Annotation version is also a practical issue. Panels designed against an older genome assembly version (e.g., GRCh37/hg19) will have coordinate offsets relative to the current GRCh38/hg38 assembly. This is operationally manageable with liftover tools, but the discrepancy must be tracked explicitly throughout the bioinformatics pipeline to avoid misalignment of probe coordinates against the analysis reference.
Probe Length and Tiling Strategy
Probe length in hybridization capture applications is typically 80 to 120 nt. Within this range, longer probes offer higher Tm and greater thermodynamic stability of the probe-target duplex, which translates to better capture efficiency at high stringency wash conditions. Shorter probes within this range provide more design flexibility in regions where longer probe placement is constrained by GC content or repeat proximity.
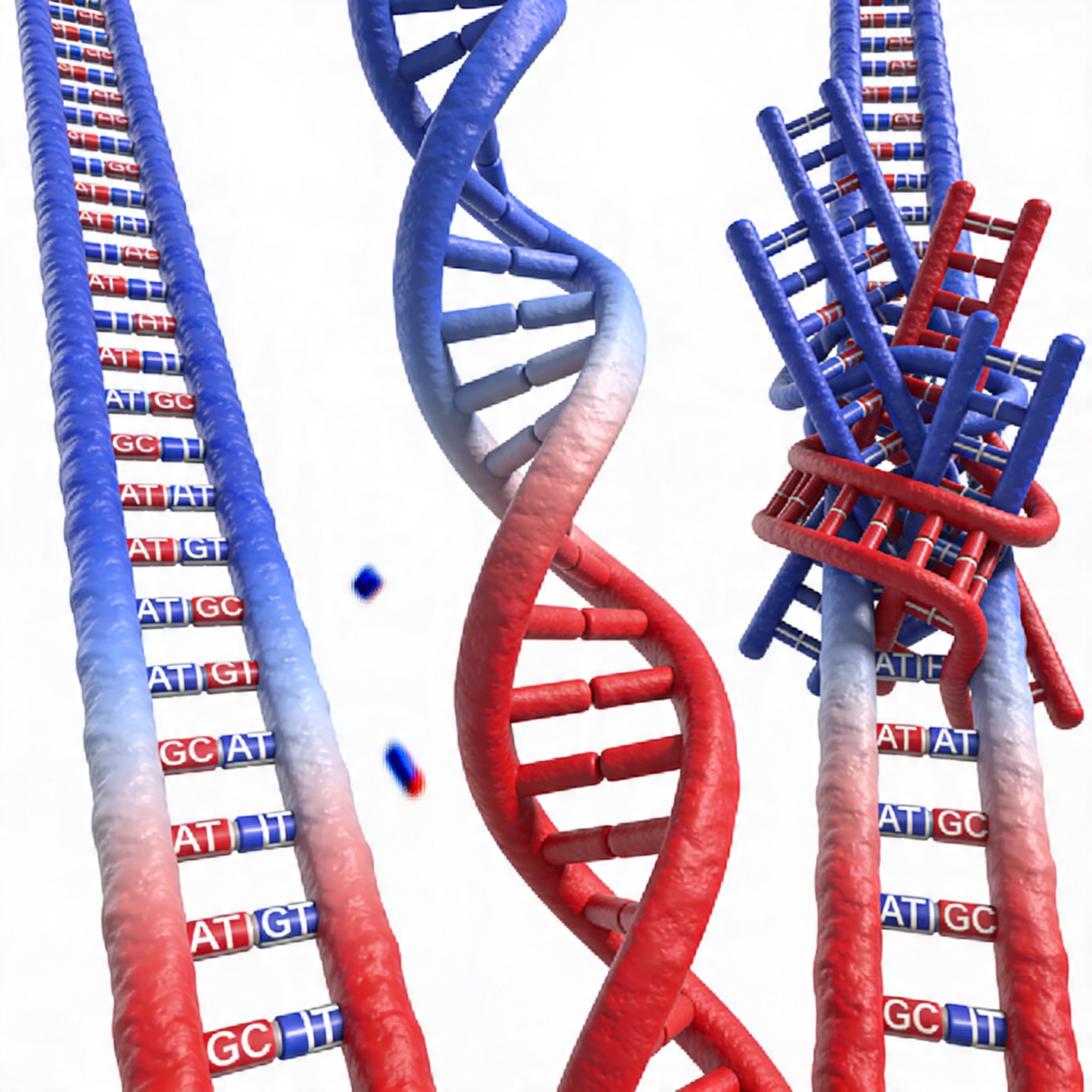

Tiling strategy — the pattern in which probes are distributed across each target region — determines both coverage uniformity and total probe count. Three main tiling patterns are used in practice:
- End-to-end (contiguous) tiling: Adjacent probes abut without overlap. This achieves the most efficient probe utilization per target base but leaves no redundancy for positions where a single probe performs poorly due to local sequence context.
- Step tiling with defined overlap: Adjacent probes overlap by a fixed number of bases (commonly 20 to 60 nt). This increases redundancy at each target base and improves uniformity, at the cost of increasing probe count.
- Boosted tiling: Selective increase in probe density at specific positions known to capture poorly — high-GC exons, AT-rich loci, or regions adjacent to repetitive elements. Boosting adds probes only where needed rather than uniformly increasing density across the entire panel.
The boosting approach is operationally more efficient than uniform high-density tiling because it concentrates the additional probe count where it is most needed. Dynegene applies AI-driven probe design and boosting to identify problematic loci and apply compensatory probe density adjustments, which is the technical basis for its claim of improved bait-to-target ratio and reduced sequencing burden relative to simpler tiling approaches.
GC Content Management
GC content is the most consequential single sequence property affecting probe hybridization performance in targeted sequencing applications. This is because the melting temperature of a DNA duplex increases with GC content (each G:C base pair contributes approximately 4°C to Tm versus 2°C for each A:T pair). Probes with very high GC content have Tm values substantially above the hybridization temperature, causing them to adopt stable intramolecular secondary structures (hairpins, G-quadruplexes) that compete with probe-target hybridization. Probes with very low GC content have Tm values below the hybridization temperature, causing them to hybridize inefficiently or dissociate during wash steps.
In practice, probe GC content below approximately 25% or above approximately 75% is associated with significantly reduced capture efficiency. The consequence in sequencing data is coverage dropout — regions where the read depth is insufficient for reliable variant calling, even when total sequencing output is adequate.
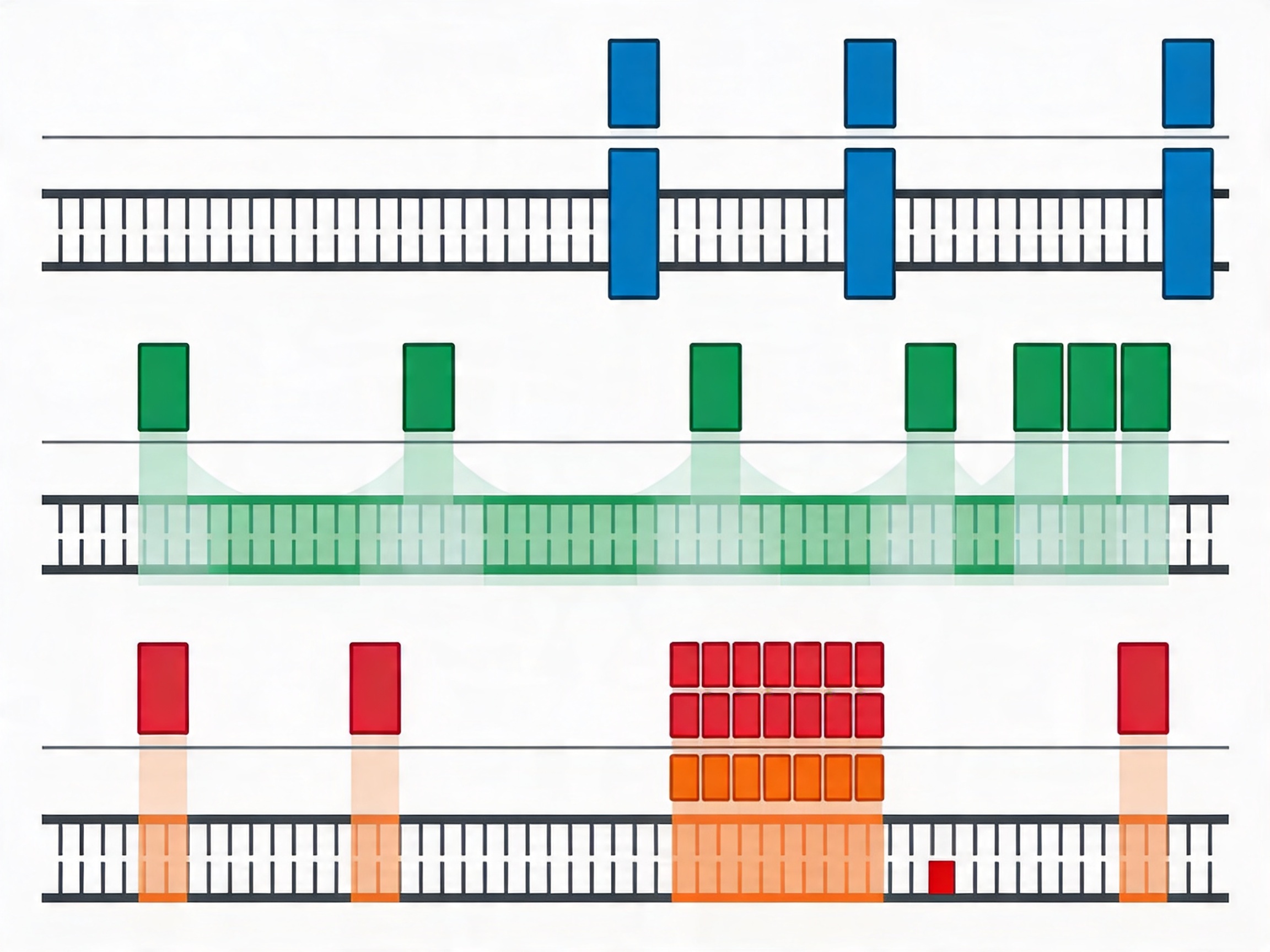
Design strategies for GC-extreme regions:
- Increased tiling density (boosting): Adding more probes at high-GC loci compensates for the reduced efficiency of each individual probe, increasing the probability that at least some probes achieve productive hybridization.
- Alternate strand probing: In cases where the sense strand probe is severely GC-biased, designing the antisense probe may provide a more balanced GC content at the same genomic position.
- Modified probe chemistry: Locked nucleic acid (LNA) modifications or other backbone modifications can increase duplex stability at GC-reduced probe sequences, effectively raising Tm without changing the sequence. This is a more expensive option and is typically reserved for specific problematic loci rather than applied broadly.
- dsDNA probes for GC-extreme regions: Double-stranded DNA probes produced by nick translation or random priming have been shown to perform better than ssDNA probes for GC-rich targets in some hybridization formats, which is part of the rationale for Dynegene's hybrid probe strategy combining dsDNA and ssDNA probe types using single-base parallel amplification labeling technology.
Repeat Element Masking
Repetitive elements in the human genome — SINEs (Alu, B1, B2), LINEs (L1, L2), DNA transposons, low-complexity sequences, and satellite repeats — account for approximately 45–50% of the human genome sequence. Probes that overlap with repetitive elements hybridize to every copy of that repeat element in the genome, not just the copy at the intended target locus. The result is capture of non-specific library fragments from throughout the genome, which manifests as reduced on-target rate and inflated sequencing cost on off-target reads.
The standard in silico approach is to apply RepeatMasker against the relevant reference genome before finalizing probe coordinates. Any probe position with significant overlap against an annotated repeat element should be replaced with a nearby non-repeat position, or handled with a modified design strategy if no clean alternative probe position exists within the target region.
Two nuances are operationally important:
- Partially repeat-overlapping probes: A probe that overlaps a repeat element at only its terminal 10 to 15 nt may still provide locus-specific capture, particularly if the unique non-repeat portion is long enough to dominate the hybridization specificity. This should be evaluated on a case-by-case basis, and validation sequencing is advisable before deploying such probes in a production panel.
- Pseudogene-adjacent regions: Probes designed against a gene of interest may also hybridize efficiently to a pseudogene elsewhere in the genome if the sequence similarity is high enough. RepeatMasker does not identify pseudogene similarity; this requires a separate BLAST-based analysis during design QC.
Probe Chemistry: ssDNA Probes, dsDNA Probes, and the Case for Hybrid Strategies
Single-stranded DNA (ssDNA) probes synthesized by chip-based oligo pool synthesis are the standard probe format for hybridization capture. They are produced by high-throughput parallel synthesis, are amenable to 5' or 3' biotin labeling during synthesis, and have well-characterized hybridization kinetics under standard capture conditions.
The limitation of ssDNA probes is at GC-extreme sequences and in applications requiring very high sensitivity. For these contexts, double-stranded DNA (dsDNA) probes produced by nick translation or random priming of a pool template can provide improved capture performance. The mechanism is different — dsDNA probes capture targets by strand displacement hybridization rather than simple complement hybridization — and the density of biotin incorporation during nick translation labeling provides more consistent pulldown efficiency than terminal-only labeling of ssDNA probes.
Biotinylation: Position, Density, and Quality Control
Biotinylation is the chemical linkage that allows probe-target duplexes to be captured on streptavidin magnetic beads. Three aspects of biotinylation affect panel performance:
Label position: Biotin at the 5' or 3' terminus is the standard for ssDNA probes synthesized on chip platforms. Terminal biotinylation is produced during synthesis by incorporating a biotinylated phosphoramidite at the defined terminus. For dsDNA probes produced by nick translation, biotin is incorporated throughout the probe body as biotinylated dNTPs are substituted for natural dNTPs during the labeling reaction — this distributed labeling provides multiple biotin-streptavidin attachment points per probe molecule, which contributes to robust pulldown efficiency.
Label density: For ssDNA probes, one biotin per probe molecule is standard. For nick translation-labeled dsDNA probes, biotin density is controlled by the ratio of biotinylated to natural dNTPs in the labeling reaction. Very high biotin density can increase steric hindrance during hybridization; very low density reduces pulldown efficiency. Optimized labeling conditions for dsDNA probes are validated against capture efficiency benchmarks.
Quality control: Probes with insufficient or absent biotinylation will not be pulled down efficiently regardless of hybridization performance. QC of biotinylation is typically performed by HABA assay (4'-hydroxyazobenzene-2-carboxylic acid) or by functional capture efficiency testing using a reference library. Poor biotinylation quality is a common cause of unexpectedly low on-target rates that is unrelated to probe design.
Custom Probes vs. Pre-Built Commercial Panels
Commercially available hybridization capture panels — whole exome kits, pan-cancer gene panels, HRD panels — are validated products with fixed content, established protocols, and documented performance specifications. For programs where the target gene set aligns with the commercial panel content, a validated commercial panel eliminates the design and validation work required for a custom panel.
However, fixed panel content is a fundamental constraint. A panel designed for broad pan-cancer sequencing covers hundreds of cancer-relevant genes at a fixed tiling density. A program focused on a specific therapeutic target landscape, a rare disease gene cohort, a specific population with a distinct variant spectrum, or a non-human pathogen genome cannot use a pan-cancer panel and obtain meaningful results.
Dynegene's custom probe service addresses this constraint directly. The NGS custom probes service supports both fully custom panel design from user-provided target coordinates and spike-in augmentation of standard panel content — for example, extending the QuarStar Pan-Cancer panels with additional targets specific to the program's therapeutic focus.
For programs that need whole exome coverage as the baseline plus additional custom targets, Dynegene's Whole Exome Sequencing Probes provide the exome backbone with spike-in capability for extended content.
Custom probe design is specifically more appropriate than a commercial panel when:
- The target list includes disease-specific loci, novel biomarkers, or therapeutic target genes not present in available commercial panels
- Non-coding regulatory regions, intron-exon boundaries for splice variant detection, or structural variant breakpoint regions must be covered
- The application involves a non-human genome (pathogen, model organism, agricultural species) for which no validated commercial panel is available
- Sample volume is large enough that the lower per-sample reagent cost of a custom panel offsets the upfront design and synthesis investment
- The research question requires control over panel content — for example, including internal calibration sequences, synthetic spike-in targets, or population-specific variant hotspots
Sample Type Considerations in Probe Design
The sample type from which the DNA library is prepared affects which probe design parameters matter most. Standard probe design guidelines are validated for high-quality genomic DNA extracted from fresh tissue or cell lines. FFPE-derived DNA and cell-free DNA introduce specific challenges that should inform probe design choices.
FFPE-Derived DNA
Formalin fixation cross-links DNA to proteins and introduces chemical modifications including cytosine deamination (C→U transitions) and strand breaks. FFPE DNA libraries have shorter fragment size distributions and higher rates of sequencing artifacts than gDNA libraries. For capture panels used with FFPE samples, tiling density should be increased relative to the gDNA default to compensate for the reduced capture efficiency from shorter, more damaged fragments. Probe chemistry considerations also apply: dsDNA probes can be less effective with highly fragmented FFPE libraries due to the shorter available complementary target length. Dynegene's DNA Library Preparation Kit is specifically designed for FFPE, cfDNA, and gDNA inputs and should be used upstream of capture to ensure library quality compatible with the probe panel's design specifications.
Cell-Free DNA (cfDNA)
cfDNA from plasma is typically 160 to 200 bp in length (corresponding to mononucleosomal fragments) with a low-input mass per sample. Capture panels used with cfDNA applications — circulating tumor DNA detection, prenatal testing, transplant monitoring — require high capture efficiency at low target mass. Probe designs for cfDNA should use shorter tiling steps than for gDNA to ensure that even the shortest cfDNA fragments have sufficient probe overlap for efficient capture. On-target rate requirements for cfDNA applications are higher than for gDNA applications because sequencing depth is the primary determinant of variant detection sensitivity at low allele fractions.
Validation Metrics for Custom Panels
Validation before large-scale deployment confirms that the designed panel performs as specified in the actual assay conditions.
Standard validation metrics and their interpretation:
|
Metric
|
Definition
|
Benchmark
|
|
On-target rate
|
% sequenced reads mapping to probe-targeted regions
|
≥80% for research panels; ≥90% for clinical-grade panels; Dynegene WES panels achieve ≥98%
|
|
Uniformity (0.2× mean)
|
% target bases covered at ≥20% of mean depth
|
≥95% is typical for high-quality panels
|
|
Coverage breadth
|
% target bases at minimum depth (e.g., 20×, 100×, 500×)
|
Depends on application; variant calling at <5% VAF typically requires ≥500× mean depth
|
|
Duplicate rate
|
% PCR duplicate reads
|
Should be monitored; high duplicate rates reduce effective coverage, especially for cfDNA
|
|
GC bias
|
Coverage fold-change across GC deciles
|
Ideally <2-fold across the full GC range
|
|
Off-target enrichment
|
Fold-enrichment over background for non-targeted regions
|
Higher is better; a panel with 98% on-target rate achieves ~50-fold enrichment over whole-genome background for a typical exome-scale target
|
For oncology panels targeting somatic variants at low allele fractions, sensitivity and specificity for variant detection should be assessed using reference standards (e.g., NIST reference samples, Horizon Discovery reference controls) in addition to standard bioinformatics QC metrics.
Application Areas for Custom NGS Capture Probes
Hybridization capture panels serve a wide range of NGS applications. The probe design priorities differ by application:
- Oncology targeted panels: High on-target rate and uniformity are critical for reliable somatic variant calling. Custom panels cover cancer driver genes, gene fusion breakpoints, HRD genomic signatures (QuarXeq HRD Panel), and HLA loci. Dynegene's commercial pan-cancer panels (QuarStar Pan-Cancer Lite Panel 3.0, QuarStar Liquid Pan-Cancer Panel 3.0) cover a broad range of cancer-relevant genes and can be extended with custom spike-in content.
- Rare disease and Mendelian genetics: Coverage of canonical coding exons, splice donor/acceptor sites, and known pathogenic intronic variants. Panels may also require coverage of deep intronic regions where splicing-altering variants have been characterized.
- Whole exome sequencing: Comprehensive coverage of all protein-coding exons. Dynegene's QuarXeq Whole Exome Sequencing Probes cover >99% of CCDS, RefSeq, and GENCODE-annotated genes with ≥98% on-target rate.
- Infectious disease and metagenomics: Custom probes for pathogen detection or enrichment from clinical or environmental samples. Dynegene's PathoSeq 450 Pathogen Library provides a broad-spectrum pathogen panel; custom probe design extends this to specific organisms or outbreak-relevant variants.
- Epigenomics: Targeted methylation sequencing using probes designed against CpG-dense regions. Dynegene's QuarStar Human MethylCap Panel addresses ultra-large panel coverage for human methylome research.
- Mitochondrial genomics: Complete mitochondrial genome coverage for rare disease diagnosis and population genetics. Dynegene's QuarXeq Mitochondrial Probes provide dedicated mitochondrial capture capability.
Complete Hybridization Capture Workflow at Dynegene
Custom probe design does not operate in isolation. The full workflow from DNA input to sequencing-ready captured library requires validated reagents at each step. Dynegene supports the complete workflow:
- Library preparation: DNA Library Preparation Kit compatible with gDNA, cfDNA, and FFPE inputs; Fragmentation Reagent for enzymatic fragmentation; NGS adapters (U-adapters and Y-adapters) for library construction.
- Hybridization capture: Custom probes from the NGS custom probe service paired with QuarHyb Super DNA Reagent Kit hybridization buffer and Streptavidin magnetic beads for pulldown.
- Blocking: Dynegene Blocker Family for adapter sequence blocking during hybridization.
Information Required to Initiate Custom Probe Design
When submitting a custom panel project to Dynegene, providing the following information upfront enables an accurate design proposal and avoids revision cycles:
- Target region coordinates in BED format, referenced to a specified genome assembly version (GRCh38/hg38, GRCh37/hg19, or other)
- Any known problematic regions — high-GC exons, repeat-adjacent loci, pseudogene-similar sequences — that require special design attention
- Sample type and input DNA quality category (high-quality gDNA, FFPE, cfDNA, low-input)
- Intended downstream application and minimum acceptable on-target rate and uniformity specifications
- Sequencing platform and read length (affects optimal fragment size distribution target for library preparation)
- Whether spike-in augmentation of a standard Dynegene panel is preferred over a fully custom design
- Expected sample throughput (for panels that will be used at scale, this informs the cost efficiency calculation between custom synthesis and commercial panel licensing)
For technical consultation before submitting a design request, Dynegene's team is available through the Contact page. Published performance data for Dynegene's commercial and custom probe products is available in the Citation and References section, including a peer-reviewed study from Shanghai Jiaotong University directly comparing capture probe performance across multiple platforms using whole exome sequencing data.
 NGSHybridization Capture Probe NGS custom probes MRD custom probes QuarStar Human MethylCap Panel QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 QuarStar Pan-Cancer Panel 1000-Gene QuarStar HLA Panel QuarXeq HRD panel QuarXeq Mitochondrial Probes Whole Exome Sequencing Probes QuarStar Human All Exon Probes 4.0 (Heredity) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 1.0 (RNA) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Dynegene Adapter Family Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus3 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Blocker Family QuarHyb DNA Reagent Kit Pro(Methyl) Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System
NGSHybridization Capture Probe NGS custom probes MRD custom probes QuarStar Human MethylCap Panel QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 QuarStar Pan-Cancer Panel 1000-Gene QuarStar HLA Panel QuarXeq HRD panel QuarXeq Mitochondrial Probes Whole Exome Sequencing Probes QuarStar Human All Exon Probes 4.0 (Heredity) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 1.0 (RNA) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Dynegene Adapter Family Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus3 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Blocker Family QuarHyb DNA Reagent Kit Pro(Methyl) Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System Primers and Probes
Primers and Probes RNA SynthesissgRNA miRNA siRNA
RNA SynthesissgRNA miRNA siRNA



 Gene
Gene Oligo Pools
Oligo Pools CRISPR sgRNA Library
CRISPR sgRNA Library Antibody Library
Antibody Library Variant Library
Variant Library





 Tel: 400-017-9077
Tel: 400-017-9077 Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai
Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai Email:
Email: Tel: 400-017-9077
Tel: 400-017-9077 Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai
Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai Email:
Email: 







