- The Latest and Most Comprehensive Ultra-Large Panel for Human Methylome Research
The QuarStar Human MethylCap Panel (NX5001) systematically covers CpG islands, CpG shores, CpG shelves, promoter regions (±2 kb), enhancer regions, differentially methylated regions (DMRs), epigenome-wide association study (EWAS) loci, non-coding RNA regions (lncRNA/miRNA), disease-associated GWAS regions, and highly conserved regions (high phyloP scores).
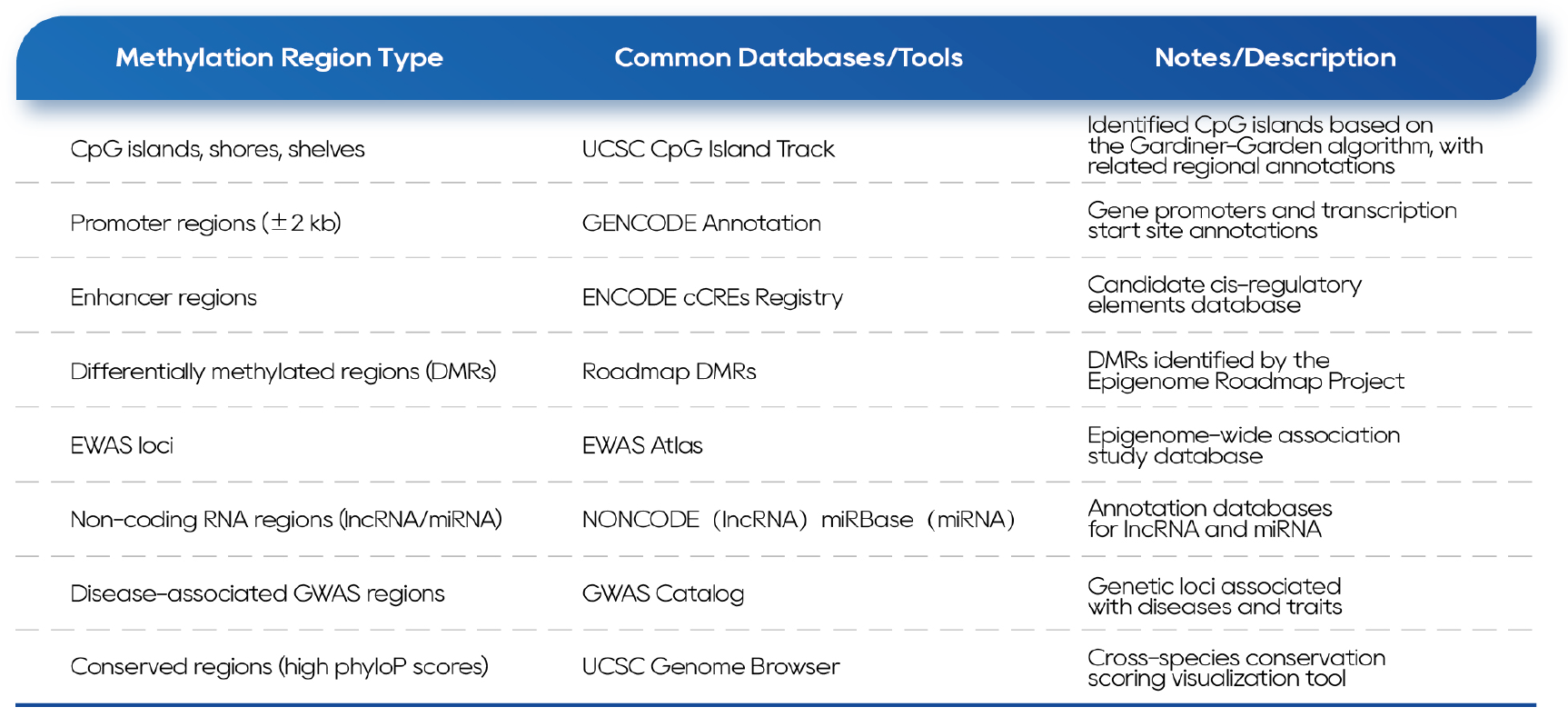
All target sites were selected based on the latest database annotations (Table 1).
Table 1. Reference Databases for the QuarStar Human MethylCap Panel
The QuarStar Human MethylCap Panel covers 96.65–99.42% of the loci included in multiple mainstream domestic and international large methylation panels and methylation array products (Figure 1-1). In addition, nearly one million high-value CpG sites have been newly incorporated, placing this panel at a leading level in terms of coverage breadth and annotation completeness (Figure 1-2, Figure 1-3).
Figure 1. Comprehensive Coverage of the QuarStar Human MethylCap Panel
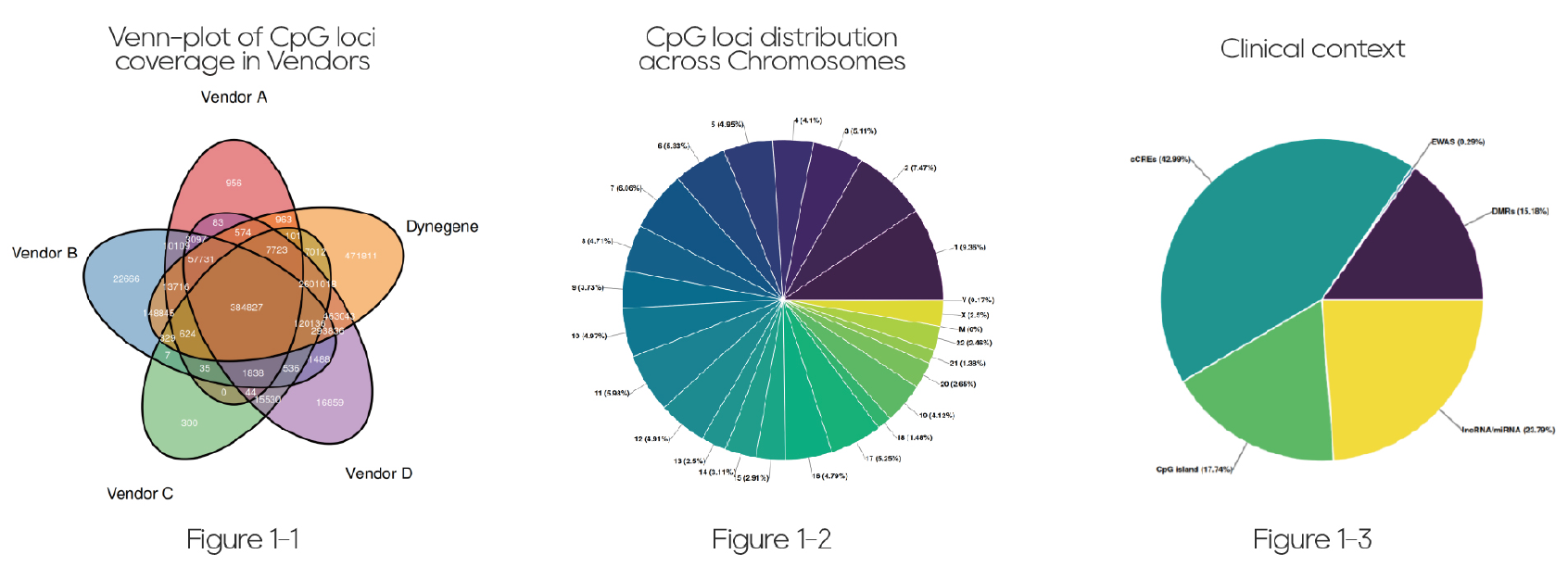
Figure 1-1. Overlap between the QuarStar Human MethylCap Panel and mainstream methylation array products and large methylation panels (Vendor A, Vendor B, Vendor C, Vendor D).
Figure 1-2. Chromosomal distribution of CpG sites in the QuarStar Human MethylCap Panel.
Figure 1-3. Regional distribution of CpG sites in the QuarStar Human MethylCap Panel (measured in base pairs).
- Degenerate Base Probe Design Reduces Cost and Improves Coverage Efficiency
Building upon the conventional “four-strand” probe design strategy, Dynegene introduces a unique degenerate base (N/R/S/Y) synthesis approach. This strategy enables the synthesis of multiple combinatorial probes at a single synthesis position, significantly reducing synthesis costs (Figure 2).
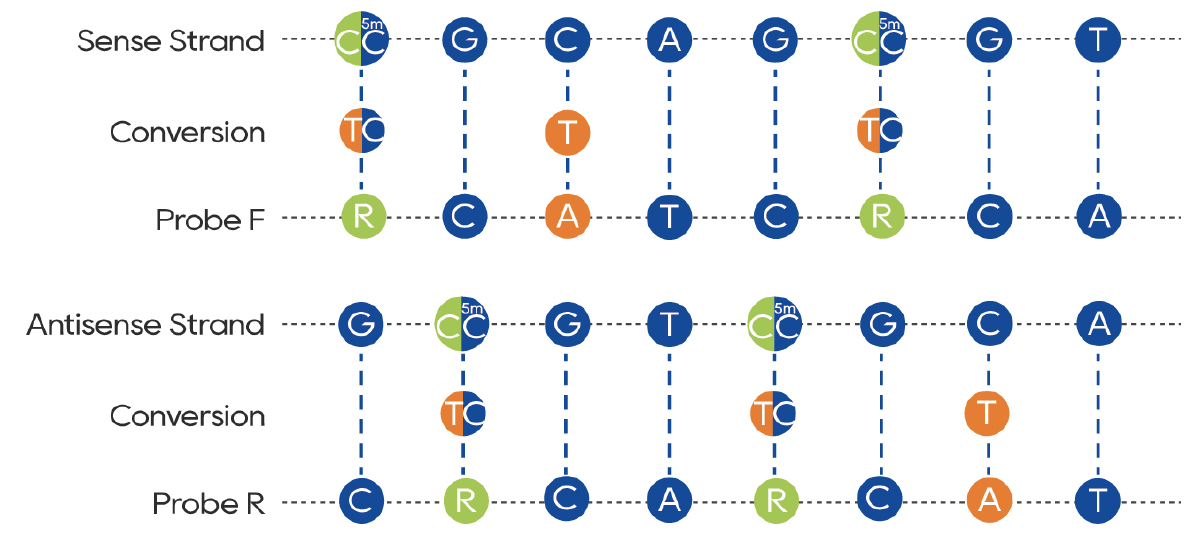
Figure 2. Principles of Methylation Probe Design:
For A, T, and G sites, the reverse complementary sequence is directly used.
For C at non-CpG sites, A is designed in the probe.
For C at CpG sites, which may be either methylated or unmethylated, R (A/G) is incorporated in the probe design.
- Methylation-Specific Blocking Reagent: QuarHyb DNA Methyl Enhancer
To address the reduced sequence complexity and decreased GC content of methylation libraries, Dynegene has developed a methylation capture enhancer — QuarHyb DNA Methyl Enhancer (NF5010).
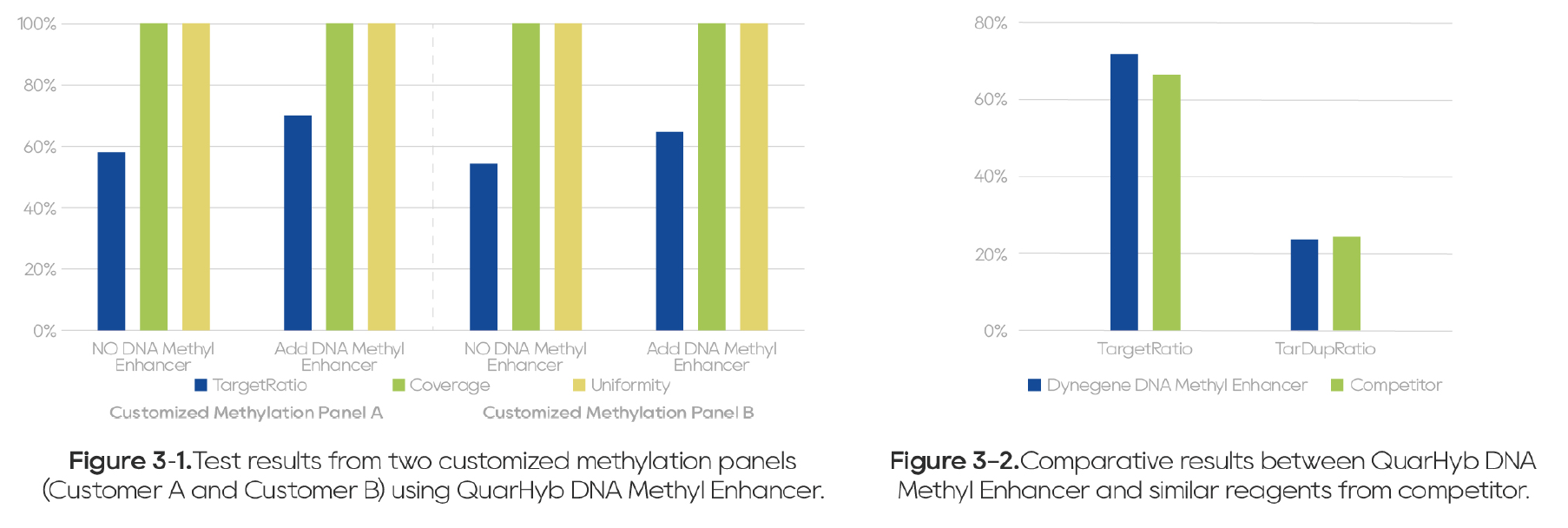
Without affecting key performance indicators such as Coverage and Uniformity, this reagent improves blocking efficiency and significantly reduces off-target rates (Figure 3).
Figure 3. Superior Blocking Performance of QuarHyb DNA Methyl Enhancer
- Excellent Capture and Detection Performance
The QuarStar Human MethylCap Panel demonstrates outstanding core performance metrics:
Target Ratio: 93.20%, Coverage: 99.02%, Uniformity: 98.47% (Figure 4-1). The panel also shows excellent reproducibility, with a Pearson correlation coefficient of 0.99 (Figure 4-2).
Consistency analysis with WGBS shows a high correlation in methylation levels (Figure 5, R = 0.98), providing a reliable option for both academic research and exploratory clinical studies.
Figure 4. Excellent Core Performance and Strong Experimental Reproducibility
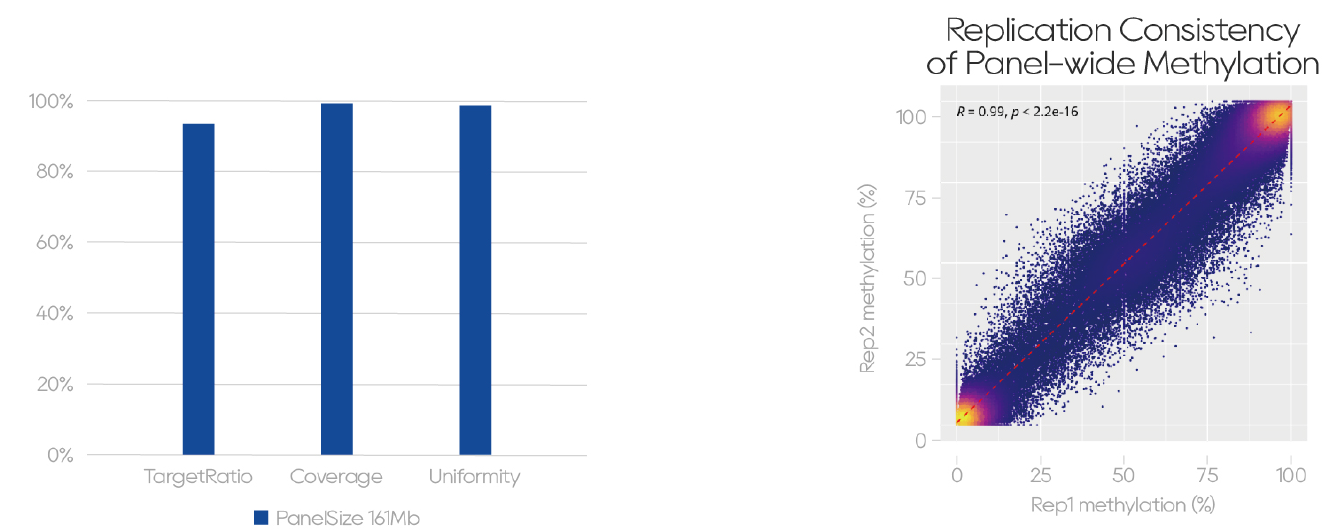
Figure 4-1. NA24385 library preparation and capture using the QuarStar Human MethylCap Panel and Dynegene methylation-specific hybridization capture reagents.
Figure 4-2. Two replicate experiments using 100 ng NA24385 from library preparation to capture, with sequencing data consistency analysis.
Figure 5. High Concordance with WGBS Methylation Levels
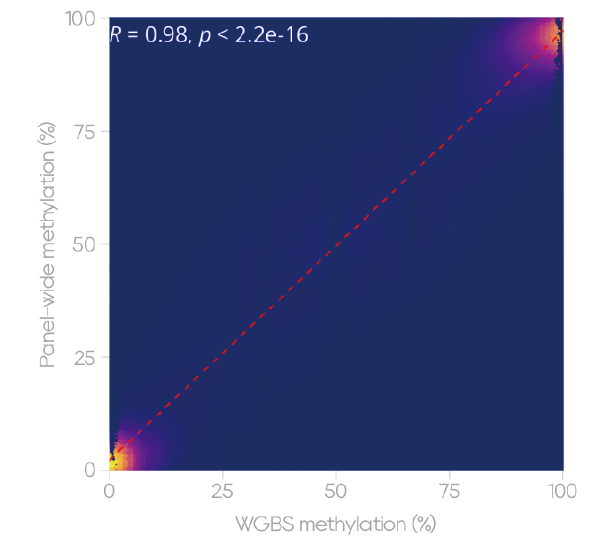
100ng NA24385 DNA was prepared into libraries and subjected to both WGBS and capture using the Dynegene QuarStar Human MethylCap Panel. Overlapping CpG sites were selected for concordance analysis.
 NGSHybridization Capture Probe NGS custom probes MRD custom probes QuarStar Human MethylCap Panel QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 QuarStar Pan-Cancer Panel 1000-Gene QuarStar HLA Panel QuarXeq HRD panel QuarXeq Mitochondrial Probes Whole Exome Sequencing Probes QuarStar Human All Exon Probes 4.0 (Heredity) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 1.0 (RNA) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Dynegene Adapter Family Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus3 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Blocker Family QuarHyb DNA Reagent Kit Pro(Methyl) Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System
NGSHybridization Capture Probe NGS custom probes MRD custom probes QuarStar Human MethylCap Panel QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 QuarStar Pan-Cancer Panel 1000-Gene QuarStar HLA Panel QuarXeq HRD panel QuarXeq Mitochondrial Probes Whole Exome Sequencing Probes QuarStar Human All Exon Probes 4.0 (Heredity) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 1.0 (RNA) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Dynegene Adapter Family Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus3 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Blocker Family QuarHyb DNA Reagent Kit Pro(Methyl) Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System Primers and Probes
Primers and Probes RNA SynthesissgRNA miRNA siRNA
RNA SynthesissgRNA miRNA siRNA



 Gene
Gene Oligo Pools
Oligo Pools CRISPR sgRNA Library
CRISPR sgRNA Library Antibody Library
Antibody Library Variant Library
Variant Library









 Tel: 400-017-9077
Tel: 400-017-9077 Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai
Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai Email:
Email: 







