Whole exome sequencing (WES) has established itself as the gold standard for identifying disease-causing coding variants in clinical genetics and oncology. At the core of every WES workflow are exome capture probes — the engineered nucleic acid sequences that selectively enrich the ~1.5% of the human genome that encodes proteins.
The choice of WES probe panel determines everything downstream: which exonic regions are captured, how uniformly they are sequenced, how much data is needed to reach clinically actionable depth, and which variant types can be reliably detected. This guide provides a comprehensive overview of WES probe technology, compares Dynegene's three exome probe panels, and maps each to its optimal clinical application.
What Is Whole Exome Sequencing?
The Exome: 1.5% of the Genome, ~85% of Disease-Causing Mutations
The human genome contains approximately 3 billion base pairs, but only about 1.5% — roughly 30 million bases — encodes proteins. This protein-coding fraction is called the exome. Despite its small size, the exome harbors an estimated 85% of disease-causing mutations identified to date. WES focuses sequencing resources on this information-dense fraction, making it far more cost-effective than whole genome sequencing (WGS) for most clinical applications.
WES vs WGS: A Cost-Effectiveness Comparison
| Factor |
WES |
WGS |
| Target size |
~35–46 Mb (exome) |
~3,000 Mb (whole genome) |
| Sequencing data needed |
10–15G (100–200x depth) |
90–120G (30x depth) |
| Cost per sample |
Lower |
Higher |
| Coding variant detection |
✓ Excellent |
✓ Excellent |
| Structural variants |
Limited |
✓ Comprehensive |
| Non-coding variants |
✗ Not covered |
✓ Covered |
| Turnaround time |
Faster (less data) |
Slower (more data) |
| Best for |
Genetic disease, tumor profiling |
Complete genomic characterization |
WES is the preferred approach when your primary objective is to identify mutations in protein-coding genes — the regions most likely to cause genetic disease, drive cancer, or predict drug response.
How WES Probes Work: The Hybridization Capture Workflow
Step 1: Library Preparation
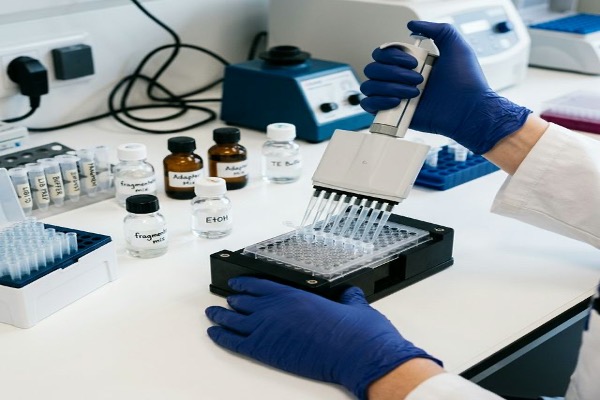

Genomic DNA is fragmented (enzymatically or mechanically) and ligated with sequencing adapters using a DNA Library Preparation Kit. Dynegene's Fragmentation Reagent provides enzymatic fragmentation for consistent fragment size distribution, and the QuarPro Superfast T4 DNA Ligase enables rapid adapter ligation.
Step 2: Hybridization Capture
The prepared library is denatured and hybridized with biotinylated WES probes. The probes bind to their complementary exonic target sequences, forming probe-target duplexes. Dynegene supports three hybridization protocols:
- Ultra-fast hybridization: As quick as 30 minutes
- Rapid hybridization: 1 hour — balanced performance and throughput
- Overnight hybridization: 12–16 hours — maximum capture efficiency
Step 3: Target Enrichment
Streptavidin magnetic beads capture the biotinylated probe-target complexes. Off-target sequences are removed through stringent washes, leaving only the enriched exomic library.
Step 4: Post-Capture Amplification and Sequencing
The enriched library undergoes PCR amplification, quality control, and is sequenced on an NGS platform. Dynegene's QuarStar 4.0 panels have been validated on the MGI sequencing platform (T7), among others.
Dynegene's WES Probe Portfolio: Three Panels for Every Application
Best for: Genetic disease testing, population genetics, baseline WES
| Specification |
Detail |
| Probe chemistry |
Double-stranded DNA |
| Genome coverage |
~35.5 Mb |
| Database design |
Refseq / CCDS / GENCODE |
| Data requirement |
~10G for ~100x effective depth |
| Hybridization |
Rapid (1 hr) and overnight; single-plex and multiplex |
| Customization |
Spike-in probes for additional targets |
| Product page |
QuarStar 4.0 Standard |
The QuarStar Standard is Dynegene's core WES panel. Key advantages:
- Excellent performance — Outstanding on-target rate and uniformity
- Cost advantage — Requires less sequencing data to achieve the same target depth
- Flexible combination — Spike-in sub-panels adapt to different testing requirements; up to 4.35 million oligos per chip enables rapid customization
- Broad compatibility — Rapid and overnight hybridization; supports multiplexed sample hybridization
Best for: Clinical oncology, precision cancer diagnosis, comprehensive tumor profiling
| Specification |
Detail |
| Probe chemistry |
Double-stranded DNA |
| Genome coverage |
~46 Mb |
| Catalog number |
NX3009 |
| Additional targets |
1,000+ tumor genes, fusions, MSI, HRD (23,157 SNPs), HLA |
| Design technology |
AI algorithm-based probe design + Boosting strategies |
| Hybridization |
Ultra-fast (30 min), rapid (1 hr), overnight |
| Compatible reagents |
QuarHyb Super DNA (NC1009, rapid), QuarHyb DNA Plus 3 (NC1018, overnight) |
| Product page |
QuarStar 4.0 Tumor |
The Tumor version's defining innovation is differential sequencing depth:
| Region |
Average Effective Depth |
Clinical Significance |
| General gene exons |
>150x (~200x) |
Germline variant detection |
| Tumor-related genes (1,000+) |
>450x (~500x) |
Somatic mutation detection |
| Hotspot regions |
>650x (~900x) |
Low-frequency driver mutations |
Built-in Oncology Biomarker Detection:
- Gene Fusions — ALK ~900x, ROS1 ~800x, RET ~1,000x, NTRK1 ~1,000x; detection validated to 2.08% AF (ddPCR-confirmed)
- HRD — 23,157 WGS-verified, GC-balanced heterozygous SNPs for LOH/TAI/LST genomic scar detection
- MSI — Integrated microsatellite markers for MSI-H/MSS determination
- HLA — Coverage for immunotherapy-relevant typing
- TMB — Comprehensive exonic coverage for accurate TMB calculation
Best for: Maximum sensitivity WES, FFPE samples, genetic disease + precision oncology
| Specification |
Detail |
| Probe chemistry |
Double-stranded RNA |
| Database design |
Refseq / CCDS / GENCODE |
| Special coverage |
TERT promoter, challenging genomic regions |
| Hybridization |
Rapid and overnight; single-plex and multiple-plex |
| Product page |
QuarXeq 3.0 |
The QuarXeq 3.0 exploits the thermodynamic advantage of RNA-RNA hybrids (stronger than RNA-DNA or DNA-DNA):
- Enhanced sensitivity — Superior target capture from degraded FFPE or low-input materials
- Dual-strand capture — dsRNA probe technology captures both sense and antisense strands simultaneously
- TERT promoter coverage — Explicitly designed to include this clinically significant challenging region
Choosing the Right WES Panel: Decision Matrix
| Your Need |
Recommended Panel |
Why |
| Genetic disease screening (germline) |
QuarStar Human All Exon Probes 4.0 (Heredity) |
Cost-effective, more comprehensive coverage, more stable performance, ~41.05 Mb, ~250x |
| Comprehensive tumor profiling (somatic) |
QuarStar 4.0 Tumor |
Differential depth, 1,000+ tumor genes, fusions, HRD, MSI |
| High-throughput clinical lab (max speed) |
QuarStar 4.0 Tumor |
Ultra-fast 30-min hybridization |
| Immunotherapy biomarkers (TMB, HLA, MSI) |
QuarStar 4.0 Tumor |
Integrated TMB + HLA + MSI in one capture |
| Carrier screening / population studies |
QuarStar 4.0 Standard |
Economical, high uniformity |
Clinical Applications in Detail
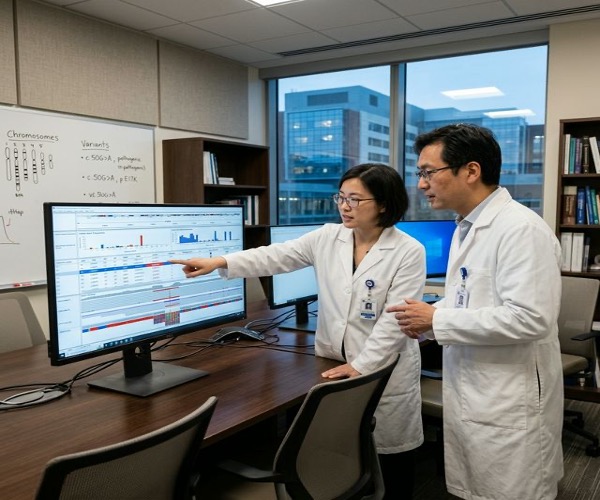

Genetic Disease Diagnostics
WES is the first-line genetic test for undiagnosed rare diseases, achieving diagnostic yields of 25–40% in clinical practice. The QuarStar Standard panel provides the most cost-effective approach for germline WES with comprehensive CDS coverage.
Precision Oncology
The QuarStar Tumor panel enables a single-capture, multi-biomarker approach:
- Somatic mutation detection in 1,000+ cancer genes at >450x depth
- Hotspot variant calling at >650x for treatment-selecting mutations
- Gene fusion detection with validated intronic coverage for ALK, ROS1, RET, NTRK1
- TMB + MSI + HRD + HLA — all from one WES capture
This integrated approach replaces multiple separate tests, reducing total cost, turnaround time, and sample consumption.
Applications in Molecular Diagnostics
Dynegene's WES probes support the full spectrum of molecular diagnostic applications: HRD, MRD, MSI, and TMB.
Complete WES Workflow Products
Hybridization Reagent Kits
Library Preparation and Accessories
Frequently Asked Questions
What are WES probes and how do they work?
WES probes are biotinylated nucleic acid sequences that selectively capture protein-coding exons (~1.5% of the genome, harboring ~85% of disease-causing mutations). During hybridization, probes bind to target exonic sequences in a DNA library; complexes are pulled down with streptavidin magnetic beads, enriching the exome for sequencing. Dynegene offers DNA-based (QuarStar, ~35.5–46 Mb) and RNA-based (QuarXeq) WES panels.
What is the difference between WES and WGS?
WES captures only protein-coding exons (10–15G data for 100–200x depth), while WGS sequences the entire genome (90–120G for 30x). WES is more cost-effective for coding variant detection — the primary goal in genetic disease and tumor profiling. WGS is preferred when structural variants or non-coding regions are needed.
What WES probe panels does Dynegene offer?
Three panels: QuarStar Standard (DNA, ~35.5 Mb, genetic disease testing and precision cancer diagnosis), QuarStar Tumor (DNA, ~46 Mb, NX3009, AI-designed, differential depth, integrated oncology biomarkers), QuarStar Heredity (DNA, ~41.05 Mb, ~250x, cost-effective with more comprehensive coverage and more stable performance), and QuarXeq 3.0 (RNA, TERT promoter, enhanced sensitivity). All support rapid and overnight hybridization with single-plex and multiplex workflows.
 NGSHybridization Capture Probe NGS custom probes MRD custom probes QuarStar Human MethylCap Panel QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 QuarStar Pan-Cancer Panel 1000-Gene QuarStar HLA Panel QuarXeq HRD panel QuarXeq Mitochondrial Probes Whole Exome Sequencing Probes QuarStar Human All Exon Probes 4.0 (Heredity) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 1.0 (RNA) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Dynegene Adapter Family Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus3 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Blocker Family QuarHyb DNA Reagent Kit Pro(Methyl) Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System
NGSHybridization Capture Probe NGS custom probes MRD custom probes QuarStar Human MethylCap Panel QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 QuarStar Pan-Cancer Panel 1000-Gene QuarStar HLA Panel QuarXeq HRD panel QuarXeq Mitochondrial Probes Whole Exome Sequencing Probes QuarStar Human All Exon Probes 4.0 (Heredity) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 1.0 (RNA) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Dynegene Adapter Family Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus3 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Blocker Family QuarHyb DNA Reagent Kit Pro(Methyl) Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System Primers and Probes
Primers and Probes RNA SynthesissgRNA miRNA siRNA
RNA SynthesissgRNA miRNA siRNA



 Gene
Gene Oligo Pools
Oligo Pools CRISPR sgRNA Library
CRISPR sgRNA Library Antibody Library
Antibody Library Variant Library
Variant Library




 Tel: 400-017-9077
Tel: 400-017-9077 Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai
Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai Email:
Email: Tel: 400-017-9077
Tel: 400-017-9077 Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai
Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai Email:
Email: 







