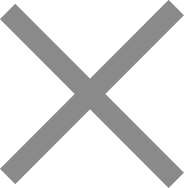
Custom oligo pools — collections of thousands to millions of oligonucleotide sequences synthesized in a single batch — are the essential building blocks for an expanding range of genomic applications. From CRISPR sgRNA screening libraries to targeted sequencing panels and antibody display libraries, oligo pool quality directly determines experimental success.
This guide covers the technology behind high-throughput oligo pool synthesis, details Dynegene's product specifications, and provides a practical framework for selecting the right oligo pool for your application.
What Is an Oligo Pool?
An oligo pool is a complex mixture of oligonucleotides, where each sequence is individually designed but synthesized together in a single reaction. Unlike conventional oligo synthesis — where each sequence is produced in a separate column — oligo pool synthesis uses chip-based parallel synthesis to produce hundreds of thousands to millions of unique sequences simultaneously.
This parallel approach dramatically reduces per-sequence cost and enables applications that require high sequence diversity, such as genome-wide CRISPR screens, variant library construction, and hybridization-based targeted sequencing.
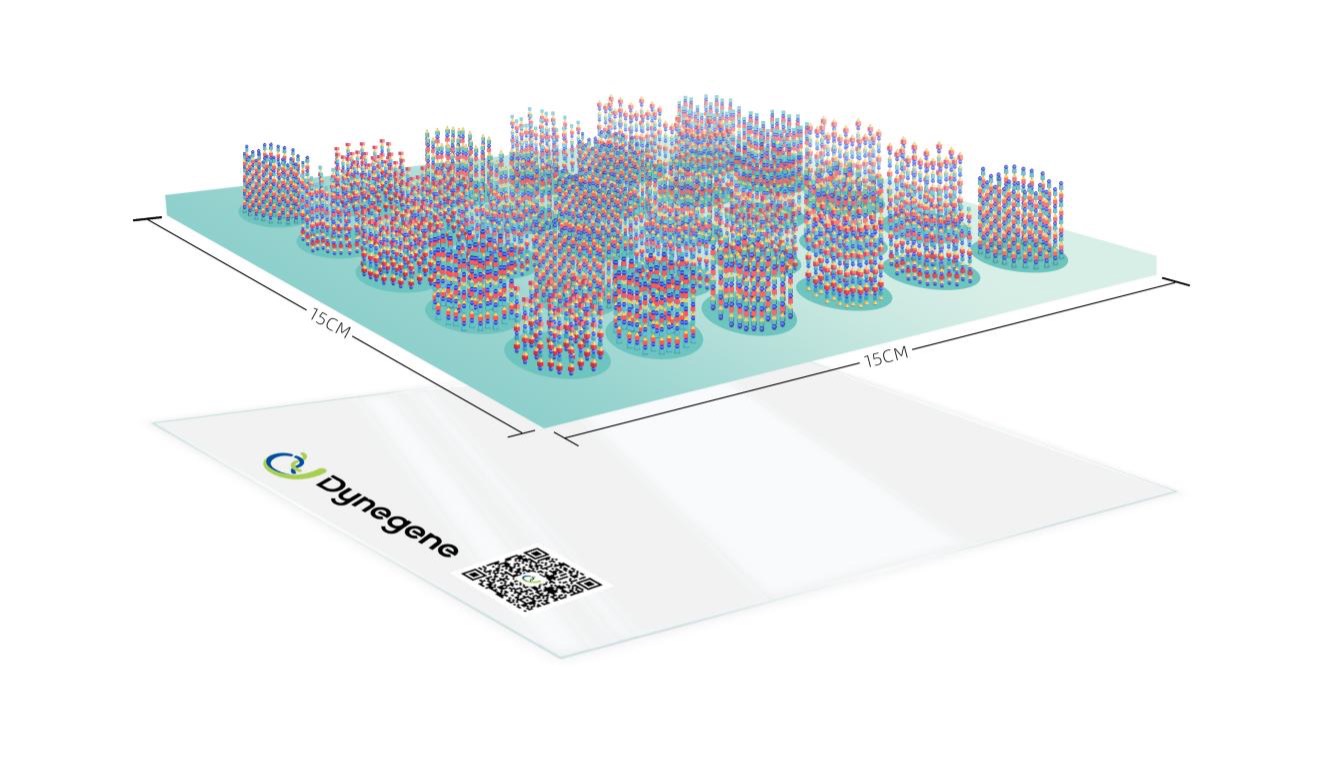
Dynegene's proprietary DYHOW (Dynegene High-throughput Oligo Warehouse) platform represents a fundamental advancement in oligo pool synthesis. The platform combines two core technologies:
3D Inkjet Printing Technology
Rather than synthesizing one oligo at a time, the DYHOW platform uses precision 3D inkjet printing to deposit nucleotide reagents onto a semiconductor chip with millions of individually addressable reaction sites. Each site synthesizes a unique oligonucleotide sequence in parallel.
Semiconductor Chip Synthesis
The chip architecture enables up to 4.35 million oligonucleotide sequences to be synthesized on a single chip, with each chip being manufactured fresh — Dynegene does not re-use chips, ensuring consistent quality across every order.
Key platform capabilities:
- Ultra-high throughput: Up to 4.35 million oligos per chip
- Length capacity: Up to 350 nt
- Accuracy: Error rate approximately 1/1500
- Uniformity: >99% sequence coverage
- Fresh chips: Every chip is newly manufactured — no re-used chips
Product Specifications
Standard Oligo Pools
| Parameter |
Specification |
| Oligo length |
Up to 350 nt |
| Pool size |
Hundreds to millions of unique sequences |
| Coverage |
>99% |
| Error rate |
~1/1500 |
| Synthesis capacity |
Up to 4.35 million oligos per chip |
| Delivery format |
Lyophilized or in solution |
| Standard turnaround |
2–3 weeks |
| Priority turnaround |
1 business day |
Ordering Information
For detailed ordering specifications, yield options, and pricing, visit the Oligo Pools ordering page.
Applications
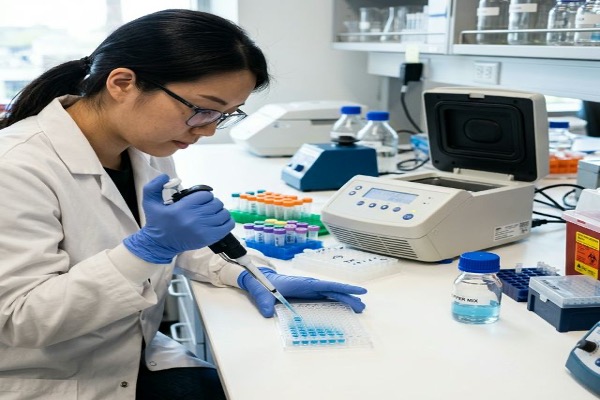

Dynegene oligo pools support a broad range of genomic and synthetic biology applications:
CRISPR sgRNA Libraries
Oligo pools are the starting material for genome-wide CRISPR sgRNA libraries. Each oligo encodes a unique guide RNA sequence, and the pool is cloned into a lentiviral vector backbone for functional screening. Dynegene's high uniformity (>99% coverage) is critical for ensuring every gene in the genome is adequately represented in the final library.
Targeted Sequencing Panels
Custom oligo pools serve as the probe source for hybridization capture panels. Researchers can design pools targeting specific gene sets, exonic regions, or variant hotspots, then biotinylate the oligos for use as capture probes in NGS workflows.
Antibody Display Libraries
Antibody library construction requires highly diverse oligo pools encoding CDR variants. The DYHOW platform's capacity for millions of unique sequences per chip makes it ideal for building high-complexity scFv or Fab libraries for phage, yeast, or ribosome display.
Variant Libraries
Site-saturation mutagenesis and variant library construction use custom oligo pools to introduce defined mutations at specific positions. Applications include enzyme engineering, protein stability studies, and deep mutational scanning (DMS).
Additional Applications
- Multiplex PCR primer panels — Hundreds of primer pairs synthesized as a single pool
- DNA data storage — Encoding digital information into synthetic DNA oligo pools
- Molecular barcoding — Unique barcodes for single-cell or spatial transcriptomics
Design Considerations for Oligo Pool Projects
1. Sequence Length
- Short oligos (20–60 nt): Primers, CRISPR guides, molecular barcodes
- Medium oligos (60–170 nt): Gene assembly fragments, capture probes
- Long oligos (170–350 nt): Full construct assembly, complex library elements
2. Pool Complexity
- Low complexity (100–10K): Targeted panels, focused CRISPR screens
- Medium complexity (10K–100K): Genome-wide CRISPR, medium antibody libraries
- High complexity (100K–4.35M): Ultra-diverse antibody libraries, comprehensive variant scanning
3. Quality Requirements
- Standard coverage (>99%): Sufficient for most screening applications
- Uniformity: Critical for CRISPR screens where under-represented guides create false negatives
Why Dynegene for Oligo Pools
Dynegene (Shanghai Dynegene Technologies) was founded as a pioneer in next-generation DNA synthesis in China and ranks among the top synthesis technology companies globally. Key differentiators:
- Proprietary DYHOW platform with independent intellectual property
- 4.35M sequences per chip — among the highest capacity in the industry
- Fresh chips for every order — no re-used substrates
- In-house synthesis to delivery — vertically integrated quality control
- Global service through authorized distributors and direct support
- Academic collaborations with leading research institutions including Shanghai Jiao Tong University
For technical questions, visit the FAQ page or explore the Technical Resources library.
Frequently Asked Questions
What is the maximum number of oligos in a single pool?
Dynegene's DYHOW platform supports up to 4.35 million unique oligonucleotide sequences per chip, with lengths up to 350 nt. The platform achieves >99% representation coverage with an error rate of approximately 1/1500. Every chip is newly manufactured — Dynegene does not re-use chips.
What applications are oligo pools used for?
Oligo pools are used for CRISPR sgRNA library construction, targeted sequencing capture probes, antibody display libraries (phage, yeast, ribosome display), variant library construction for protein engineering, multiplex PCR primer panels, molecular barcoding, and DNA data storage.
What is the turnaround time for custom oligo pools?
Standard turnaround for custom oligo pools is 2–3 weeks from order confirmation. Priority (expedited) service is available for time-sensitive projects. Contact Dynegene directly at info2@dynegene.com or through the online inquiry form for custom timeline requests.
 NGSHybridization Capture Probe NGS custom probes QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 HRD panel Whole Exome Sequencing Probes 人全外显子组探针1.0(RNA) QuarStar Human All Exon Probes 4.0 (Heredity Version) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus 2 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Adapter Family Dynegene Blocker Family Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System
NGSHybridization Capture Probe NGS custom probes QuarStar Liquid Pan-Cancer Panel 3.0 QuarStar Pan-Cancer Lite Panel 3.0 QuarStar Pan-Cancer Fusion Panel 1.0 QuarStar Pan Cancer Panel 1.0 HRD panel Whole Exome Sequencing Probes 人全外显子组探针1.0(RNA) QuarStar Human All Exon Probes 4.0 (Heredity Version) QuarStar Human All Exon Probes 4.0 (Tumor) QuarStar Human All Exon Probes 4.0 (Standard) QuarXeq Human All Exon Probes 3.0 (RNA) Library Preparation DNA Library Preparation Kit Fragmentation Reagent mRNA Capture Kit rRNA Depletion Kit QuarPro Superfast T4 DNA Ligase Hybridization Capture QuarHyb Super DNA Reagent Kit QuarHyb DNA Plus 2 Reagent Kit QuarHyb DNA Reagent Kit Plus QuarHyb One Reagent Kit QuarHyb Super Reagent Kit Pro Dynegene Adapter Family Dynegene Blocker Family Multiplex PCR QuarMultiple BRCA Amplicon QuarMultiple PCR Capture Kit 2.0 PathoSeq 450 Pathogen Library Corollary Reagent Streptavidin magnetic beads Equipment and Software The iQuars50 NGS Prep System Primers and Probes
Primers and Probes RNA SynthesissgRNA miRNA siRNA
RNA SynthesissgRNA miRNA siRNA



 Gene
Gene Oligo Pools
Oligo Pools CRISPR sgRNA Library
CRISPR sgRNA Library Antibody Library
Antibody Library Variant Library
Variant Library




 Tel: 400-017-9077
Tel: 400-017-9077 Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai
Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai Email:
Email: Tel: 400-017-9077
Tel: 400-017-9077 Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai
Address: Floor 2, Building 5, No. 248 Guanghua Road, Minhang District, Shanghai Email:
Email: 







